Listen to the Science Friday podcast where Adam Phillippy and Kateryna Makova discuss the challenges of sequencing the Y chromosome, and what doing so might mean for medical research.
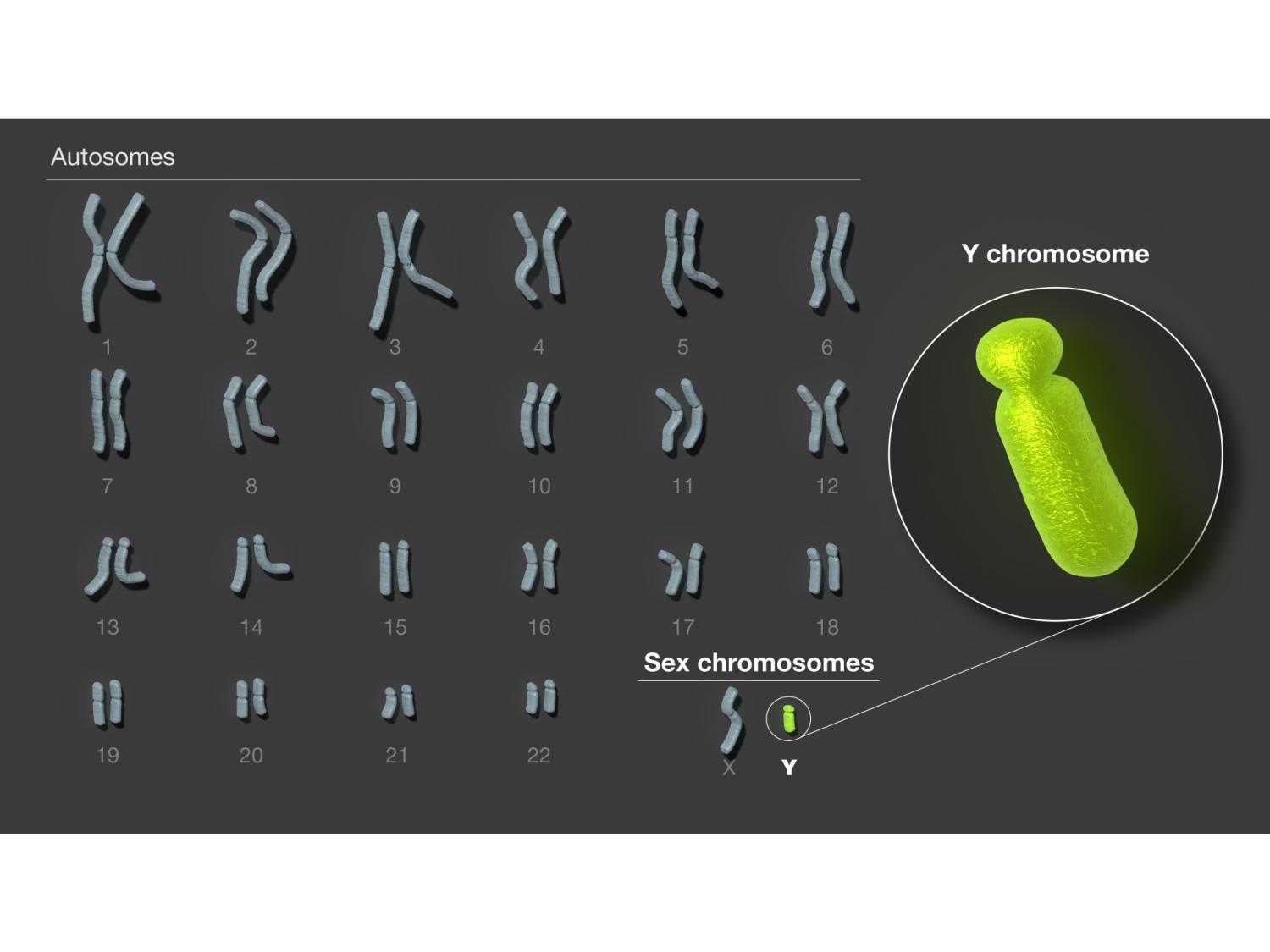
Researchers determined the full sequence of the human Y chromosome for the first time. Humans have 23 pairs of chromosomes, and Y is the smallest and last to be assembled. Credit: Provided by the National Institutes of Health. All Rights Reserved.
UNIVERSITY PARK, Pa. — The first full sequence of the last human chromosome — the Y chromosome — to be assembled is complete, thanks to an international collaboration that includes Penn State researchers. The new sequence, which fills in gaps across more than 50% of the Y chromosome’s length, uncovers genomic features with implications for fertility, such as factors in sperm production, as well as cancer risk and severity.
The Y chromosome, along with the X chromosome, is often discussed for its role in sexual development. While these chromosomes play a central role, the factors involved in human sexual development are spread across the genome and are very complex, giving rise to the array of human sex characteristics found among male, female and intersex individuals. These categories are not equivalent to gender, which is a social category. Additionally, recent work demonstrates that genes on the Y chromosome contribute to other aspects of human biology, such as cancer risk and severity, according to the collaboration.
The study, led by the Telomere-to-Telomere (T2T) Consortium, a team funded by the National Human Genome Research Institute (NHGRI), part of the National Institutes of Health, was published today (Aug. 23) in Nature.
Kateryna Makova, the Verne M. Willaman Chair of Life Sciences, professor of biology and director of the Center for Medical Genomics, led Penn State’s contributions to the study.
“The Y chromosome is by far the most difficult human chromosome to sequence and assemble,” Makova said, explaining it is highly repetitive and is present in only one copy. “Deciphering its complete sequence is a major scientific milestone. My group has been working on the Y chromosome for over 20 years, and I did not think it would be possible to obtain its complete sequence in the near future.”
Makova’s team provided Y-chromosome-specific expertise to aid in the assembly and analysis of the chromosome. Specifically, they annotated the Y chromosome’s haplogroup — the parental ancestry — and compared this assembly to a previous incomplete assembly of the chromosome. They also validated the new Y chromosome sequence with experiments in the lab and studied the distribution of non-canonical DNA — in which the nucleotide bases pair up differently than in the classic structure described by James Watson and Francis Crick — on the Y chromosome.
“The complete sequence of the Y chromosome will provide scientists and physicians with critically missing information that will assist with diagnosing and treating important genetic disorders,” Makova said. She also noted that the Y chromosome harbors several genes important for sperm production, and deletion of these genes has been associated with infertility disorders in males.
When researchers drafted the first human genome sequence 20 years ago, gaps were left in the sequences of all 24 chromosomes. However, unlike the small gaps sprinkled across the rest of the genome sequence — gaps that the T2T Consortium filled in last year — more than half of the Y chromosome’s sequence remained a mystery.
All chromosomes have some repetitive regions, but the Y chromosome is unusually repetitive, making its sequence particularly difficult to complete. Assembling sequencing data is like trying to read a long book cut into strips. If all the lines in the book are unique, it’s easier to determine the order the lines go in. However, if the same sentence is repeated thousands or millions of times, the original order of the strips is far less clear. While all human chromosomes contain repeats, about 30 million letters of the Y chromosome are repetitive sequences. It’s as if the same few sentences were repeated for half the length of the book.
To tackle the most repetitive pieces of the human genome, the T2T Consortium applied new DNA sequencing technologies and sequence assembly methods, as well as knowledge gained from generating the first gapless sequences for the other 23 human chromosomes.
“The biggest surprise was how organized the repeats are,” said Adam Phillippy, a senior investigator at NHGRI and leader of the consortium. “We didn’t know what exactly made up the missing sequence. It could have been very chaotic, but instead, nearly half of the chromosome is made of alternating blocks of two specific repeating sequences known as satellite DNA. It makes a beautiful, quilt-like pattern.”
The complete Y chromosome sequence also reveals important features of medically relevant regions. One such section of the Y chromosome is called the azoospermia factor region, a stretch of DNA containing several genes known to be involved in sperm production. With the newly completed sequence, the researchers studied the structure of a set of inverted repeats or “palindromes” in the azoospermia factor region.
“This structure is very important because occasionally these palindromes can create loops of DNA,” said Arang Rhie, NHGRI staff scientist and first author of the Nature publication. “Sometimes, these loops accidentally get cut off and create deletions in the genome.”
Deletions in the azoospermia factor region are known to disrupt sperm production, and thus these palindromes could influence fertility. With a complete Y chromosome sequence, researchers can now more precisely analyze these deletions and their effects on sperm production.
Other regions with potential medical relevance contain genes that repeat. Most genes in the human genome have two copies, one inherited from each parent. However, some genes have many copies that repeat along a stretch of DNA, sometimes referred to as a “gene array.”
The researchers focused on TSPY, another gene thought to be involved in sperm production. Copies of TSPY are organized in the second largest gene array in the human genome. Like other repetitive regions, repeating genes are challenging to analyze, so while TSPY was known to exist as many repeating copies, the specific DNA sequence and organization of this array was previously unknown. As the researchers analyzed this region, they found that different individuals contained between 10 and 40 copies of TSPY.
“When you find variation that you haven’t seen before, the hope is always that those genomic variants will be important for understanding human health,” Phillippy said. “Medically relevant genomic variants can help us design better diagnostics in the future.”
In addition to the complete Y chromosome sequence, the NHGRI-funded Human Genome Structural Variation Consortium reports the sequence of 43 diverse human Y chromosomes, also published today in the same issue of Nature. These advances complement the gapless human genome sequence released by the T2T Consortium in 2022, as well as the “pangenome” released in May of 2023 by the NHGRI-funded Human Pangenome Reference Consortium. Through these achievements, scientists have access to an abundance of new genomics resources to unravel human biology and pave the way for the future of genomic medicine.
Co-authors from Penn State on this paper include Robert Harris, assistant research professor of biology; Paul Medvedev, professor of computer science and engineering, and of biochemistry and molecular biology, and director of the Center for Computational Biology and Bioinformatics; Marta Tomaszkiewicz, assistant research professor of biomedical engineering; Allison Watwood, doctoral student in the Department of Biology; and Matthias Weissensteiner, postdoctoral research in the Department of Biology. The co-authors also included Penn State alumni — former doctoral students in the Makova Laboratory: Monika Cechova, now a postdoctoral fellow at the University of California, Santa Cruz; Arkarachai Fungtammasan, now a researcher at DNA Nexus, and Melissa Wilson, now an associate professor at Arizona State University.
A full list of authors for this paper is available here.
The NHGRI partially supported this work.
Editor’s note: The original version of this press release is available on the National Institutes of Health website.
CONTACT
Francisco Tutella, francisco@psu.edu
Work Phone: 814-863-4638