CHAPTER
9
TRANSPOSITION
OF DNA
The final method of changing the DNA in a genome that we will consider is transposition, which is the movement of DNA from one location to another. Segments of DNA with this ability to move are called transposable elements. Transposable elements were formerly thought to be found only in a few species, but now they are recognized as components of the genomes of virtually all species. In fact, transposable elements (both active and inactive) occupy approximately half the human genome and a substantially greater fraction of some plant genomes! These movable elements are ubiquitous in the biosphere, and are highly successful in propagating themselves. We now realize that some transposable elements are also viruses, for instance, some retroviruses can integrate into a host genome to form endogenous retroviruses. Indeed, some viruses may be derived from natural transposable elements and vice versa. Since viruses move between individuals, at least some transposable elements can move between genomes (between individuals) as well as within an individual’s genome. Given their prevalence in genomes, the function (if any) of transposable elements has been much discussed but is little understood. It is not even clear whether transposable elements should be considered an integral part of a species’ genome, or if they are successful parasites. They do have important effects on genes and their phenotypes, and they are the subject of intense investigation.
Transposition is related to replication, recombination and repair. The process of moving from one place to another involves a type of recombination, insertions of transposable elements can cause mutations, and some transpositions are replicative, generating a new copy while leaving the old copy intact. However, this ability to move is a unique property of transposable elements, and warrants treatment by itself.
Properties and effects of transposable elements
The defining property of transposable elements is their mobility; i.e. they are genetic elements that can move from one position to another in the genome. Beyond the common property of mobility, transposable elements show considerable diversity. Some move by DNA intermediates, and others move by RNA intermediates. Much of the mechanism of transposition is distinctive for these two classes, but all transposable elements effectively insert at staggered breaks in chromosomes. Some transposable elements move in a replicative manner, whereas others are nonreplicative, i.e. they move without making a copy of themselves.
Transposable elements are major forces in the evolution and rearrangement of genomes (Fig. 9.1). Some transposition events inactivate genes, since the coding potential or expression of a gene is disrupted by insertion of the transposable element. A classic example is the r allele (rugosus) of the gene encoding a starch branching enzyme in peas is nonfunctional due to the insertion of a transposable element. This allele causes the wrinkled pea phenotype in homozygotes originally studied by Mendel. In other cases, transposition can activate nearby genes by bringing an enhancer of transcription (within the transposable element) close enough to a gene to stimulate its expression. If the target gene is not usually expressed in a certain cell type, this activation can lead to pathology, such as activation of a proto-oncogene causing a cell to become cancerous. In other cases, no obvious phenotype results from the transposition. A particular type of transposable element can activate, inactivate or have no effect on nearby genes, depending on exactly where it inserts, it’s orientation and other factors.
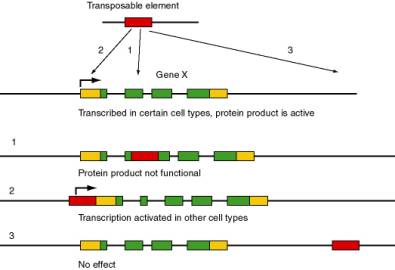
Figure 9.1. Possible effects of movement of a transposable element in the function and expression of the target gene. The transposable element is shown as a red rectangle, and the target gene (X) is composed of multiple exons. Protein coding regions of exons are green and untranslated regions are gold. The angled arrow indicates the start site for transcription.
Transposable elements can cause deletions or inversions of DNA. When transposition generates two copies of the same sequence in the same orientation, recombination can delete the DNA between them. If the two copies are in the opposite orientations, recombination will invert the DNA between them.
As part of the mechanism of transposition, additional DNA sequences can be mobilized. DNA located between two copies of a transposable element can be moved together with them when they move. In this manner, transposition can move DNA sequences that are not normally part of a transposable element to new locations. Indeed, "host" sequences can be acquired by viruses and propagated by infection of other individuals. This may be a natural means for evolving new strains of viruses. One of the most striking examples is the acquisition and modification of a proto-oncogene, such as cellular c-src, by a retrovirus to generate a modified, transforming form of the gene, called v-src. These and related observations provided insights into the progression of events that turn a normal cell into a cancerous one. They also point to the continual acquisition (and possibly deletion) of information from host genomes as a natural part of the evolution of viruses.
Parasites or symbionts?
Do the transposable elements confer some selective advantage on the "host"? Or are they merely parasitic or "selfish," existing only to increase the number of copies of the element? This critical issue is a continuing controversy. As just mentioned, certain results of transposition can be detrimental, leading to a loss of function or changes in regulation of the genes at the site of integration after movement. Also, we are starting to appreciate the intimate connection between viruses and transposable elements. Thus one can view many transposable elements as parasites on the genome. The number of transposable elements can expand rapidly in a genome. For instance, it appears that transposable elements making up a majority of the genome of maize are not abundant in the wild parent, teosinte. Thus this massive expansion has occurred since the domestication of corn, roughly within the past 10,000 years.
However, other studies indicate that the presence of transposable elements is beneficial to an organism. Two strains of bacteria, one with a normal number of transposable elements and the other with many fewer, can be grown in competitive conditions. The strain with the higher number of transposable elements has a growth advantage under these conditions. Various proposals have been made as to the nature of that advantage. One intriguing possibility is that the mechanism of transposition affords an opportunity to seal chromosome breaks. Other possible benefits have not been excluded. Thus the relationship between transposable elements and their hosts may be as much symbiotic as parasitic. Resolving these issues is an interesting challenge for future research.
Discovery of transposable elements as controlling elements in maize
The discovery of transposable elements by Barbara McClintock is a remarkable story of careful study and insightful analysis in genetics. Long before the chemical structure of genes was known, she observed that genetic determinants, called controlling elements, in maize were moving from one location to another. The controlling elements regulate the expression of other genes. The families of controlling elements are now recognized as members of the class of transposable elements that move through DNA intermediates. However, McClintock’s proposal that the controlling elements were mobile was not widely accepted for a very long time. Despite her extensive observations published in the 1930’s through the 1950’s, the interpretation that genetic elements could move was perhaps too novel. Indeed, the notion that transposable elements are active in a wide range of species as not widely accepted until the 1980’s, and new evidence continues to mount that transposable elements are more common than previously thought.
McClintock’s seminal observations relied on two complementary approaches to understanding chromosome structure and function. One was cytological, using microscopy to examine the structure of chromosomes in corn, and the other used genetics to follow the fates of the chromosomes. A full exploration of the discovery of transposable elements is the subject of excellent books. In this section, we will examine a few examples of the type of studies that were done, to give some impression of the care and insight of the work.
In essence, McClintock showed that certain crosses between maize cultivars (or strains) resulted in large numbers of mutable loci, i.e. the frequency of change at those loci is much higher than observed in other crosses. Her studies of the cultivars with mutable loci revealed a genetic element termed “Dissociation”, or Ds. Chromosome breaks occurred at the Ds locus; these could be seen cytologically, using a microscope to examine chromosome spreads from individual germ cells (sporocytes). The frequency and timing of these breaks is controlled by another locus, called “Activator” or Ac. In following crosses of the progeny, the position of Ds-mediated breaks changed, arguing that the Ds element had moved, or transposed. That was the basic argument for transposition.
Frequent chromosome breaks at Ds
The studies of Ds on chromosome 9 illustrate the combination of morphological examination of chromosome structure plus genetic analysis to show that the controlling elements were mobile in the genome. Chromosome 9 of maize has a knob at the end of its short arm, making it easy to identify when chromosome spreads are examined in the microscope. In some versions of chromosome 9, a long stretch of densely staining heterochromatin extends beyond the knob, forming a hook (but shown as a green oval in Fig. 9.2). These morphologically distinct versions of the same chromosome can have different sets of alleles for the genes on this chromosome. As diagrammed in Fig. 9.2, several genes affecting the appearance of corn kernels are on this chromosome. The colorless gene has three alleles we will consider: the recessive c allele confers no color, the C allele (dominant to c) makes the kernel colored, and the I allele, which is dominant to C, confers no color. The recessive allele sh makes the kernel look shrunken, whereas the dominant Sh is nonshrunken. The recessive bz confers a bronze phenotype, whereas the dominant Bz does not. The recessive wx gives the kernel a waxy appearance, whereas the dominant Wx makes the kernel starchy. Of course, all the phenotypes stated for recessive alleles are for the homozygous or hemizygous (only one allele present, e.g. because the other is deleted) states. Thus different versions of chromosome 9 that have a distinctive appearance in the microscope (knob or extended heterochromatin at the ends) confer different phenotypes on progeny.
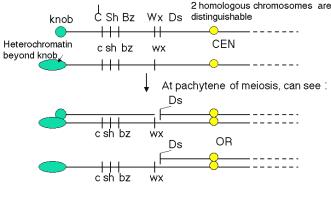
Figure 9.2. Two homologs of chromosome 9 can be distinguished both by appearance and genetic determinants. The short of chromosome 9 can have either a knob or extended heterochromatin, denoted by the green circle and the elongated oval, respectively. The alleles of each of the genes diagrammed confer different phenotypes. The yellow circle is the centromere (CEN). Ds is the dissociation element that leads to chromosome breaks.
The two homologs will pair to form a bivalent during the pachytene phase of meiosis I. Ordinarily, the two homologs will form a continuous complex with no disruptions, as shown in panel 3 and 3a of Fig. 9.3. However, when Ds is on the short arm of chromosome 9 and an Ac element is also present in the genome, a break in one of the chromosomes in the pair can be seen when spreads of chromosomes are examined in the microscope (panels 4, 4a, 5 and 5a). One can identify chromosome 9 specifically because of the knob or extended heterochromatin at its end. In panels 4 and 4a, a break has occurred in the knob chromosome (with the dominant alleles diagrammed in Fig. 9.2), leaving the other homolog intact, with the recessive alleles and marked by the extended heterochromatin). Both a break and a crossover occurred in the chromosome pair shown in panels 5 and 5a. In a given strain, the break usually occurred in the same position, so the genetic element at the site of the break was called “dissociation”, or Ds.

Figure 9.3.
Cytological examination in the microscope reveals breaks on morphologically
marked chromosomes. The figure shows photomicrographs (panels 3, 4 and 5) and
interpretative drawings (panels 3a, 4a and 5a) of paired homologous chromosomes
at the pachytene phase of meiosis. The telomere at the end of the short arm of
chromosome 9 (labeled a in the pictures) can be either a darkly staining spot,
called a knob, or an elongated hook. The centromere is labeled b and the breaks
are labeled c. These images are adapted from a 1952 paper from McClintock in
the Cold Spring Harbor Symposium on Quantitative Biology.
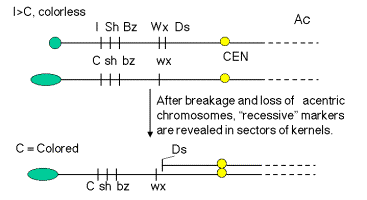
These effects of these frequent breaks in the chromosomes could be seen phenotypically when the sporocytes (e.g. pollen grains) with Ds and Ac were used to fertilize ova of a known genotype. For instance, pollen from a plant homozygous for the “top” chromosome in Fig. 9.4.A. will carry the dominant alleles (indicated by capitalized names) for all the loci shown. When this pollen is used to fertilize an ovum that has the recessive alleles along chromosome 9, the resulting corn kernel will show the phenotypes specified by the dominant alleles. However, if the chromosome with the dominant alleles also has a Ds element, and Ac is present in the genome, the chromosome will break in some of the cells making up the kernel as some stage in development. Then the region between Ds and the telomere will be lost from this chromosome, and the phenotype of the progeny cells will be determined by the recessive alleles on the other chromosome. For example, the phenotype of the kernel outlined in Fig. 9.4.A. will be colorless, nonshrunken and nonwaxy (starchy), but the sector of the kernel derived from a cell in which a break occurred at Ds will be colored, shrunken and waxy. In more detail, I is dominant to C (which itself is dominant to c; hence the capital letter). This gives a colorless seed when the chromosome is intact, but after the break, I is lost and C is left, generating a colored phenotype. Similarly, prior to the break the starch will not be waxy (Wx is dominant), but after the break one sees waxy starch because only the recessive wx allele is present.
A.
B.
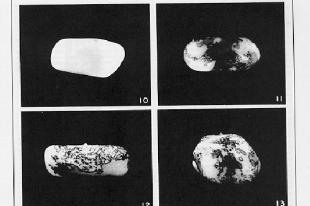
Figure 9.4. Breaks at Ds can reveal previously hidden phenotypes of recessive alleles (in the presence of Ac). A. Prior to the break, the dominant alleles along the chromosome with Ds (I Sh Bz Wx, shown at the top) determine the phenotype. [The part of the corn kernel showing the phenotypes studied is actually triploid, resulting from fertilizing a diploid ovum with a haploid pollen grain. For this discussion the diploid ovum is homozygous recessive, and only one copy is shown, C sh bz wx.] After the chromosome breaks at Ds, which occurs frequently in the presence f Ac, the phenotype will be determined by the recessive alleles thus revealed, C sh bz wx. B. Kernels with variegating color. The chromosome breaks in some but not all cells, and only those with the broken cells show the new phenotypes. All the progeny of the cells with a broken chromosome are located adjacent to each other, resulting in a patch of cells with the same new phenotype. Thus the new phenotype is variegating across the kernel. The kernel shown in panel 10 is colorless, determined by the I allele. Panels 11-13 show patches of colored kernel, representing patches of cells in which the I allele has been deleted because of the chromosome break and revealing the effect of the C allele. B was adapted from McClintock in the Cold Spring Harbor Symposium on Quantitative Biology.
These frequent breaks occurring at different times in different cells derived from the fertilized ovum can produce a sectored, patched or stippled appearance to the corn kernel, as illustrated in Fig. 9.4.B. The phenotype differs in the various parts of the kernel, even though all the cells are derived from the same parental cell (i.e. the kernel is clonal). This differing phenotype in a clonal tissue is called variegation. Each sector is the product of the expansion of one cell. When a chromosome breaks in that cell, thereby removing the effect of a dominant allele I that was making the seed colorless, then all the progeny in that sector would be colored (from the effects of the C allele in the example in Fig. 9.4.B.).
Variegating phenotypes can be caused by breaks such as those described here, but in other cases they result from modifications to the regulation of genes. Variegation is a fairly common occurrence, and is especially visible in flower petals, as illustrated for the wildflowers in Fig. 9.5.
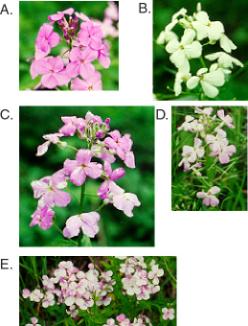

Figure 9.5. Variegation
in sectors of wildflower petals. A wildflower blooms all over the middle
Atlantic states in the USA in late May and June. My neighbors call this wild
flox. It is an invasive plant, but it is pretty when it blooms. It has two
predominant flower colors, purple (A) and white (B). However, a casual
examination of the plants reveals sectored petals at a moderate frequency
(C-E). This is a variegating phenotype of unknown origin. It can produce white
sectors on purple petals (C) or purple sectors on white petals (D, E).
Question 9.1. How does this phenotype in Fig. 9.5, panels C-E, differ from partial dominance, e.g. with the purple allele dominant and the white allele recessive?
Ds can
appear at new locations
By following several generations of a maize cultivar with Ds on chromosome 9, McClintock observed that Ds could move to new locations. As outlined in Fig. 9.6, chromosomal rearrangements associated with Ds activity can appear at several different positions on chromosome 9. If, e.g., Ds were centromeric to Wx in one generation, but it was between I and Sh in a subsequent generation, the simplest explanation is that it had moved. These observations are the basis for the notion that Ds is transposable.
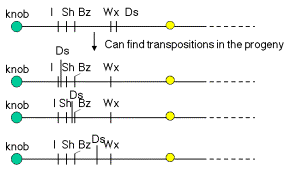
Figure 9.6. Ds activity can appear at new locations on chromosome 9.
How does one know that Ds is present at different locations on chromosome 9? The effects of breaking the chromosome (Fig. 9.2 and 9.4) depend on where Ds is. The position of the observable break (e.g. bottom panel of Fig. 9.3) and the genetic consequences in terms of which recessive allele are revealed, will differ depending on where Ds is.
Question 9.2. What phenotype in kernels would result if the second chromosome after the arrow in Fig. 9.6 were in a heteroduplex with the recessive chromosome shown in Fig. 9.4.A, and Ac were also present?
The example of a single Ds affecting all the genes telomeric to it on chromosome 9 shows a particular controlling element can simultaneously regulate the expression of genes involved in a variety of biochemical pathways. The Ac element is needed to activate the mobility of any Ds element, regardless of its chromosomal location. Thus controlling elements can operate independently of the chromosomal location of the controlling element. These observations show that the controlling element is distinct from the genes whose expression is being regulated.
The movement of Ds to new locations on chromosome 9 is associated with other types of recombinations that involve breaks, including duplications and inversions. Other types of transposable elements also cause inversions and duplications in their vicinity when they move.
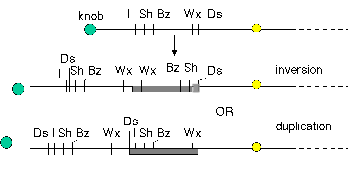
Figure 9.7. The appearance of Ds at a new location is associated with duplications and inversions. Some of the rearranged chromosomes found in progeny in which Ds had moved are shown.
Insertion of a controlling element can
generate an unstable allele of a locus
The insertion of a controlling element can generate an unstable allele of a locus, designated mutable. This instability can be seen both in somatic and in germline tissues. The instability can result from reversion of a mutation, due to the excision and transposition of the controlling element. After excision and re-integration, the transposable element can alter the expression of a gene at the new location. This new phenotype indicated that the element was mobile.
An example of the effects of integration and excision of a transposable element can be seen at the bronze locus in maize (Fig. 9.8). The aleurone is the surface layer of endosperm in a kernel of maize. The wild type has a deep bluish-purple color. This is determined by the bronze locus. The Bz allele is dominant and confers the bluish-purple color to the aleurone. The bz allele is recessive, and gives a bronze color to the aleurone when homozygous. In Bz kernels, anthocyanin is produced. Bz encodes UDPglucose:flavanoid 3-O-glucosyltransferase (UFGT), an enzyme needed for anthocyanin production. The loss-of-function bz alleles have no UFGT activity, and the bluish-purple anthocyanins are not produced. Some alleles of bronze show an unstable, or mutable, phenotype. In the bz-m alleles, clones of cells regain the bluish-purple color characteristic of Bz cells. This produces patches of bluish-purple color in the aleurone of kernels (Fig. 9.8).
This mutation in the bz-m allele is the insertion of the Ds (dissociation) transposable element. Ds disrupts the function of the UFGT gene to give a bronze color to the seed kernel. In the presence of the Ac (activator) element, the Ds can excise from the locus, restoring a functional UFGT gene (and a bluish-purple color). This occurs in some but not all cells in the developing seed and is clonally inherited, resulting in the patches of blue on a bronze background for each kernel.
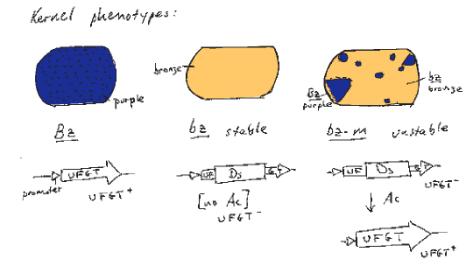
Figure 9.8. Frequent excision of a Ds allele generates an unstable, or mutable, phenotype at the bronze locus.
Current methods for observing transposition and transposable elements
Movement of DNA segments can be observed by a variety of modern techniques. In organisms with a short generation time, such as bacteria and yeast, one can simply monitor many generations for the number and positions of a family of repeated DNA elements by blot-hybridization analysis of genomic DNA. Using a Ty-1 DNA fragment as a probe, about 20 hybridizing bands could be seen at the start of an experiment, meaning that about 20 copies were present in the yeast genome. The size of the restriction fragment containing each element was distinctive, as determined by restriction endonuclease cleavage sites that flanked the different locations of each element. After growing for many generations, some new bands were observed, showing that new Ty-1 elements had been generated and moved to new locations. These observations led to this family of repeats being christened Ty-1, for transposable element, yeast, number 1.
Evidence for transposition in many organisms comes from analysis of new mutations. Transposable elements appear to be the major source of new mutation in Drosophila, and they have been shown to cause mutations in bacteria, fungi, plants and animals. One example from humans is a new mutation causing hemophilia. A patient from a family with no prior history was diagnosed with hemophilia, resulting from an absence of factor VIII. By molecular cloning techniques, Kazazian and his colleagues showed that the mutant factor VIII gene had a copy of a LINE1, or L1, repeat inserted. In contrast to most L1 repeats in the human genome, whose sequences have diverged from a predicted source gene, the sequence of this L1 was very close to that predicted for an active L1. Tests showed that the patient’s parents did not carry this mutation in their factor VIII genes. Screening a genomic library for L1s that were almost identical to the mutagenic L1 revealed a full-length, active L1 that was the source, on a different chromosome. The appearance of a new L1 in the factor VIII gene, making an allele that was not present in the parents, is a strong argument for transposition. The further studies identifying a source gene and showing that the source gene is active in transposition make the evidence unequivocal.
Now that it is recognized that most repetitive elements in many species result from transposition events, it is easy to find transposable elements or their progeny. A comprehensive database of repetitive elements in many species is maintained as RepBase (J. Jurka) and the program RepeatMasker (Green and Smit) will widely used to find matches to these repeats. RepeatMasker is available as a server on the World Wide Web, and one can find many repeats in a query sequence quickly and comprehensively. Virtually all these repeats are made by transposition.
Transposition occurs by insertion into a staggered break in a chromosome
A common property of virtually all transposable elements is that they move by inserting into a staggered break in a chromosome, i.e. one strand is slightly longer than the other at the break (Fig. 9.9). The first indication of this was the observation that the same short DNA sequence is found on each side of a transposable element. The sequence within these flanking direct repeats (FDRs) is distinctive for each copy of the transposable element, but the size of the FDR is characteristic of a particular family of transposable elements. Some families of transposable elements have FDRs as short as 4 bp and other families have FDRs as long as 12 bp. However, within a particular family, the sequence of the FDR will differ between individual copies. These FDRs are hallmarks of transposable elements.

Figure 9.9. Flanking direct repeats are generated by insertions at staggered breaks.
Since the FDRs are distinctive for each copy, they are not part of the transposable element themselves. Some families of transposable elements do have repeated sequences at their flanks that are identical for all members of the family, but these are integral parts of the transposable element. The variation in sequence of the FDRs indicates that they are generated from the target sites for the transposition events. If the transposable element inserted into a break in the chromosome that left a short overhang (one strand longer than the other), and this overhang were filled in by DNA polymerase as part of the transposition, then the sequence of that overhang would be duplicated on each side of the new copy. Such a break with an overhang is called a staggered break. The size of the staggered break would determine the size of the FDR.
Mechanistic studies of the enzymes used for transposition have shown that such staggered breaks are made at the target site prior to integration and are repaired as part of the process of transposition (see below). The staggered breaks are used in transposition both by DNA intermediates and by RNA intermediates.
Major classes of transposable elements
The two major classes of transposable elements are defined by the intermediates in the transposition process. One class moves by DNA intermediates, using transposases and DNA polymerases to catalyze transposition. The other class moves by RNA intermediates, using RNA polymerase, endonucleases and reverse transcriptase to catalyze the process. Both classes are abundant in many species, but some groups of organisms have a preponderance of one or the other. For instance, bacteria have mainly the DNA intermediate class of transposable elements, whereas the predominant transposable elements in mammalian genomes move by RNA intermediates.
Transposable elements that move via DNA intermediates
Among the most thoroughly characterized transposable elements are those that move by DNA intermediates. In bacteria, these are either short insertion sequences or longer transposons.
An insertion sequences, or IS, is a short DNA sequence that moves from one location to another. They were first recognized by the mutations they cause by inserting into bacterial genes. Different insertion sequences range in size from about 800 bp to 2000 bp. The DNA sequence of an IS has inverted repeats (about 10 to 40 bp) at its termini (Fig. 9.10.A.). Note that this is different from the FDRs, which are duplications of the target site. The inverted repeats are part of the IS element itself. The sequences of the inverted repeats at each end of the IS are very similar but not necessarily identical. Each family of insertion sequence in a species is named IS followed by a number, e.g. IS1, IS10, etc.
An insertion sequence encodes a transposase enzyme that catalyzes the transposition. The amount of transposase is well regulated and is the primary determinant of the rate of transposition.
Transposons are larger transposable elements, ranging in size from 2500 to 21,000 bp. They usually encode a drug resistance gene or other marker besides the functions required for transposition (Fig. 9.10.B.). One type of transposon, called a composite transposon, has an IS element at each end (Fig. 9.10.C.). One or both IS elements may be functional; these encode the transposition function for this class of transposons. The IS elements flank the drug resistance gene (or other selectable marker). It is likely that the composite transposon evolved when two IS elements inserted on both sides of a gene. The IS elements at the end could either move by themselves or they can recognize the ends of the closely spaced IS elements and move them together with the DNA between them. If the DNA between the IS elements confers a selective advantage when transposed, then it will become fixed in a population.
Question 9.3. What are the predictions of this model for formation of a composite transposon for the situation in which a transposon in a small circular replicon, such as a plasmid?
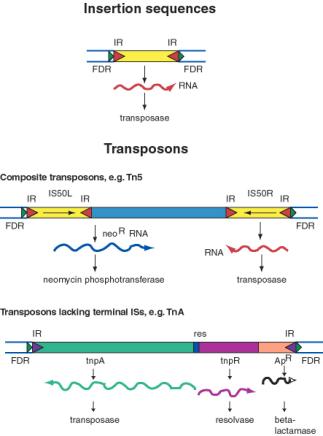
Figure 9.10. General structure of insertion sequences and transposons. Flanking direct repeats (FDRs) are shown as green triangles, inverted repeats (IRs) are red or purple triangles, insertion sequences (ISs) are yellow boxes with red triangles at the end, and other genes are boxes of different colors. The boxes and triangles include both strands of duplex DNA. DNA outside the FDRs is shown as one thick blue line for each strand. Tn5 has an IS50 element on each side, in an inverted orientation. Transcripts are shown as curly lines with an arrowhead pointing in the direction of transcription. The neoR gene for Tn5 is composed partly of the leftward IS (ISL) and partly of other sequences (included in the blue box). The transposase for Tn5 is encoded in the rightward IS (ISR).
The TnA family of transposons has been intensively studied for the mechanism of transposition. Members of the TnA family have terminal inverted repeats, but lack terminal IS elements (Fig. 9.10). The tnpA gene of the TnA transposon encodes a transposase, and the tnpR gene encodes a resolvase. TnA also has a selectable marker, ApR, which encodes a beta-lactamase and makes the bacteria resistance to ampicillin.
Transposable elements that move via DNA intermediates are not limited to bacteria, but rather they are found in many species. The P elements and copia family of repeats are examples of such transposable elements in Drosophila, as are mariner elements in mammals and the controlling elements in plants. Indeed, the general structure of controlling elements in maize is similar to that of bacterial transposons. In particular, they end in inverted repeats and encode a transposase. As illustrated in Fig. 9.11, the DNA sequences at the ends of an Ac element are very similar to those of a Ds element. However, internal regions, which normally encode the transposase, have been deleted. This is why Ds elements cannot transpose by themselves, but rather they require the presence of the intact transposon, Ac, in the cell to provide the transposase. Since transposase works in trans, the Ac element can be anywhere in the genome, but it can act on Ds elements at a variety of sites. Note that Ac is an autonomous transposon because it provides its own transposase and it has the inverted repeats needed to act as the substrate for transposase.
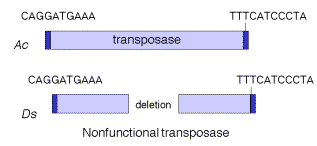
Figure 9.11. Structure of Ac and Ds controlling elements in maize is similar to that of an intact (Ac) or defective (Ds) transposon.
Mechanism of DNA-mediated transposition
Some families of transposable elements that move via a DNA intermediate do so in a replicative manner. In this case, transposition generates a new copy of the transposable element at the target site, while leaving a copy behind at the original site. A cointegrate structure is formed by fusion of the donor and recipient replicons, which is then resolved (Fig. 9.12). Other families use a nonreplicative mechanism. In this case, the original copy excises from the original site and move to a new target site, leaving the original site vacant.
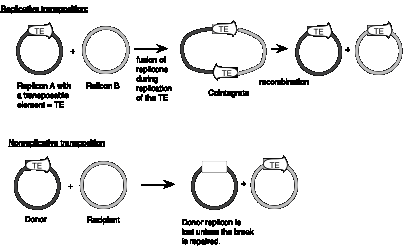
Figure 9.12. Contrasts between replicative and nonreplicative transposition. The transposable element (TE) is shown as an open arrow. The thick line for each replicon represents double stranded DNA; the different shadings represent different sequences.
Studies of bacterial transposons have shown that replicative transposition and some types of nonreplicative transposition proceed through a strand-transfer intermediate (also known as a crossover structure), in which both the donor and recipient replicons are attached to the transposable element (Fig. 9.13). For replicative transposition, DNA synthesis through the strand-transfer intermediate produces a transposable element at both the donor and target sites, forming the cointegrate intermediate. This is subsequently resolved to separate the replicons. DNA synthesis does not occur at the crossover structure in nonreplicative transposition, thus leaving a copy only at the new target site. In an alternative pathway for nonreplicative transposition, the transposon is excised by two double strand breaks, and is joined to the recipient at a staggered break (illustrated at the bottom of Fig. 9.12).
In more detail, there are two steps in common for replicative and nonreplicative transposition, generating the strand-transfer intermediate (Fig. 9.13).
(1) The transposase encoded by a transposable element makes four nicks initially. Two nicks are made at the target site, one in each strand, to generate a staggered break with 5' extensions (3' recessed). The other two nicks flank the transposon; one nick is made in one DNA strand at one end of the transposon, and the other nick is made in the other DNA strand at the other end. Since the transposon has inverted repeats at each end, these two nicks that flank the transposon are cleavages in the same sequence. Thus the transposase has a sequence-specific nicking activity. For instance, the transposase from TnA binds to a sequence of about 25 bp located within the 38 bp of inverted terminal repeat (Fig. 9.10). It nicks a single strand at each end of the transposon, as well as the target site (Fig. 9.13). Note that although the target and transposon are shown apart in the two-dimensional drawing in Fig. 9.13, they are juxtaposed during transposition.
(2) At each end of the transposon, the 3' end of one strand of the transposon is joined to the 5' extension of one strand at the target site. This ligation is also catalyzed by transposase. ATP stimulates the reaction but it can occur in the absence of ATP if the substrate is supercoiled. Ligation of the ends of the transposon to the target site generates a strand-transfer intermediate, in which the donor and recipient replicons are now joined by the transposon.
After formation of the strand-transfer intermediate, two different pathways can be followed. For replicative transposition, the 3' ends of each strand of the staggered break (originally at the target site) serve as primers for repair synthesis (Fig. 9.13). Replication followed by ligation leads to the formation of the cointegrate structure, which can then be resolved into the separate replicons, each with a copy of the transposon. The resolvase encoded by transposon TnA catalyzes the resolution of the cointegrate structure. The site for resolution (res) is located between the divergently transcribed genes for tnpA and tnpR (Fig. 9.10). TnA resolvase also negatively regulates expression of both tnpA and tnpR (itself).
For nonreplicative transposition, the strand-transfer intermediate is released by nicking at the ends of the transposon not initially nicked. Repair synthesis is limited to the gap at the flanking direct repeats, and hence only one copy of the transposon is left. This copy is ligated to the new target site, leaving a vacant site in the donor molecule.
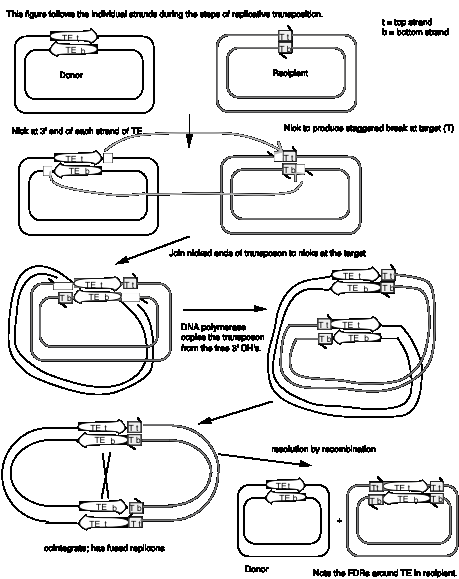
Figure 9.13. Mechanism
of transposition via a strand-transfer intermediate.
The enzyme transposase can recognize specific DNA sequences, cleave two duplex DNA molecules in four places, and ligate strands from the donor to the recipient. This enzyme has a remarkable ability to generate and manipulate the ends of DNA. A three-dimensional structure for the Tn5 transposase in complex with the ends of the Tn5 DNA has been solved by Rayment and colleagues. One static view of this protein DNA complex is in Fig. 9.14.A. The transposase is a dimer, and each double-stranded DNA molecule (donor and target) is bound by both protein subunits. This orients the transposon ends into the active sites, as shown in the figure. Also, an image with just the DNA (Fig. 9.14.B.) shows considerable distortion of the DNA helix at the ends. This recently determined structure is a good starting point to better understand the mechanism for strand cleavage and transfer.
A.
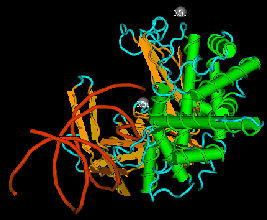 B.
B.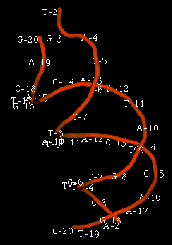
Figure 9.14. Three-dimensional structure of the Tn5 transposase in complex with Tn5 transposon DNA. A. The dimer of the Tn5 transposase is shown bound to a fragment of duplex DNA from the end of the transposon. Alpha helices are green cylinders, beta sheets are yellow-brown, flat arrows and protein loops are blue wires. The DNA is a duplex of two red wires, one for each strand. B. The DNA is shown without the protein and with the nucleotides labeled. The end of the DNA at the top of this panel is oriented into the active site in the middle of the protein in panel A. The structure was determined by Davies DR, Goryshin IY, Reznikoff WS, Rayment I. (2000) “Three-dimensional structure of the Tn5 synaptic complex transposition intermediate.” Science 289:77-85. These images was obtained by downloading the atomic coordinates from the Molecular Modeling Database at NCBI, viewing them with CN3D 3.0 and saving static views as screen shots. The file for observing a virtual three-dimensional image is available at the course website.
Transposable elements that move via RNA intermediates
Transposable DNA sequences that move by an RNA
intermediate are called retrotransposons. They are very common in eukaryotic organisms, but
some examples have also been found in bacteria.
Some retrotransposons have long terminal repeats
(LTRs) that regulate expression (Fig. 9.15). The LTRs were initially discovered
in retroviruses. They have now been seen in some but not all retrotransposons.
They have a strong promoter and enhancer, as well as signals for forming the 3’
end of mRNAs after transcription. The presence of the LTR is distinctive for
this family, and members are referred to as LTR-containing retrotransposons.
Examples include the yeast Ty-1 family and retroviral proviruses in vertebrates.
Retroviral proviruses encode a reverse transcriptase and an endonuclease, as
well as other proteins, some of which are needed for viral assembly and
structure.
Others retrotransposons are in the large and
diverse class of non-LTR retrotransposons (Fig. 9.15). One of the most
prevalent examples is the family of long interspersed repetitive elements,
or LINEs. It was initially found in mammals but has now been found in a broad
range of phyla, including fungi. The first and most common LINE family in
mammals is the LINE1 family, also called L1. An older family, but discovered
later, is called LINE2. Full-length LINEs are about 7000 bp long, and there are
about 10,000 copies in humans. Many other copies are truncated from the 5’
ends. Like retroviral proviruses, the full-length L1 encodes a reverse
transcriptase and an endonuclease, as well as other proteins. However, the
promoter is not an LTR. Other abundant non-LTR retrotransposons, initially
discovered in mammals, are short interspersed repetitive elements,
or SINEs. These are about 300 bp long. Alu repeats, with over a million copies,
comprise the predominant class of SINEs in humans. Non-LTR retrotransposons
besides LINEs are found in many other species, such as jockey repeats in Drosophila.
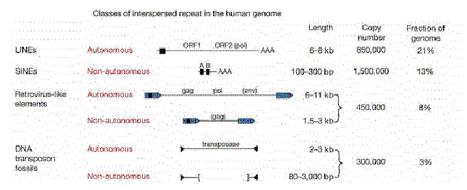
Figure 9.15. Four classes of transposable elements make up the vast majority of human repetitive DNA. From the Nature paper “Initial sequencing and analysis of the human genome,” by the International Human Genome Consortium.
Extensive studies in of genomic DNA sequences have allowed the reconstruction of the history of transposable elements in humans and other mammals. The major approach has been to classify the various types of repeats (themselves transposable elements), align the sequences and determine how different the members of a family are from each other. Since the vast majority of the repeats are no longer active in transposition, and have no other obvious function, they will accumulate mutations rapidly, at the neutral rate. Thus the sequence of more recently transposing members are more similar to the source sequence than are the members that transposed earlier. The results of this analysis show that the different families of repeats have propagated in distinct waves through evolution (Fig. 9.16). The LINE2 elements were abundant prior to the mammalian divergence, roughly 100 million years ago. Both LINE1 and Alu repeats have propagated more recently in humans. It is likely that the LINE1 elements, which encode a nuclease and a reverse transcriptase, provide functions needed for the transposition and expansion of Alu repeats. LINE1 elements have expanded in all orders of mammals, but each order has a distinctive SINE, all of which are derived from a gene transcribed by RNA polymerase III. This has led to the idea that LINE1 elements provide functions that other different transcription units use for transposition.
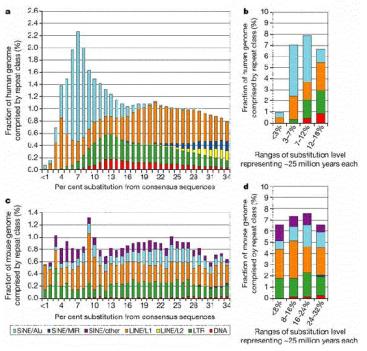
Figure 9.16. Age distribution of repeats in human and mouse. The LINE2 and MIR repeats propagated before the mammalian radiation, about 100 million years ago, but Alu repeats are formed by recent transpositions in primates (light blue portion of the bar graphs in a and b). The LINE1 and LTR repeats are transposing with about the same frequency as they have historically in the mouse lineage (panels c and d), but few repeats are still transposing in human (panels a and b). From the Nature paper “Initial sequencing and analysis of the human genome,” by the International Human Genome Consortium.
Mechanism of retrotransposition
Although the mechanism of retrotransposition is not completely understood, it is clear that at least two enzymatic activities are utilized. One is an integrase, which is an endonuclease that cleaves at the site of integration to generate a staggered break (Fig. 9.17). The other is RNA-dependent DNA polymerase, also called reverse transcriptase. These activities are encoded in some autonomous retrotransposons, including both LTR-retrotransposons such as retroviral proviruses and non-LTR-retrotransposons such as LINE1 elements.
The RNA transcript of the transposable element interacts with the site of cleavage at the DNA target site. One strand of DNA at the cleaved integration site serves as the primer for reverse transcriptase. This DNA polymerase then copies the RNA into DNA. That cDNA copy of the retrotransposon must be converted to a double stranded product and inserted at a staggered break at the target site. The enzymes required for joining the reverse transcript (first strand of the new copy) to the other end of the staggered break and for second strand synthesis have not yet been established. Perhaps some cellular DNA repair functions are used.
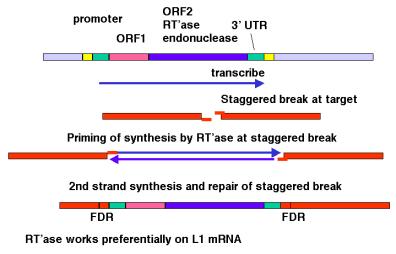
Figure 9.17. Transposition via an RNA-intermediate in retrotransposons. LINE1, or L1 repeats are shown as an example.
The
model shown in Fig. 9.17 is consistent with any RNA serving as the template for
synthesis of the cDNA from the staggered break. However, LINE1 mRNA is clearly
used much more often than other RNAs. The basis for the preference of the
retrotransposition machinery for LINE1 mRNA is still being studied. Perhaps the
endonuclease and reverse transcriptase stay associated with the mRNA that
encodes them after translation has been completed, so that they act in cis with respect to the LINE1 mRNA. Other repeats that
have expanded recently, such as Alu
repeats in humans, may share sequence determinants with LINE1 mRNA for this cis preference.
Clear evidence that retrotransposons can move via an RNA intermediate came from studies of the yeast Ty-1 elements by Gerald Fink and his colleagues. They placed a particular Ty-1 element, called TyH3 under control of a GAL promoter, so that its transcription (and transposition) could be induced by adding galactose to the media. They also marked TyH3 with an intron. After inducing transcription of TyH3, additional copies were found at new locations in the yeast strain. When these were examined structurally, it was discovered that the intron had been removed. If the RNA transcript is the intermediate in moving the Ty-1 element, it is subject to splicing and the intron can be removed. Hence, these results fit the prediction of an RNA-mediated transposition. They demonstrate that during transposition, the flow of Ty-1 sequence information is from DNA to RNA to DNA.
Question 9.4. If yeast Ty-1 moved by the mechanism illustrated for DNA-mediated replicative transposition in Fig. 9.13, what would be predicted in the experiment just outlined? Also, would you expect an increase in transposition when transcription is induced?
Additional consequences of transposition
Not only can transposable elements interrupt genes or disrupt their regulation, but they can cause additional rearrangements in the genome. Homologous recombination can occur between any two nearly identical sequences. Thus when transposition makes a new copy of a transposable element, the two copies are now potential substrates for recombination. The outcome of recombination depends on the orientation of the two transposable elements relative to each other. Recombination between two transposable elements in the same orientation on the same chromosome leads to a deletion, whereas it results in an inversion if they are in opposite orientations (Fig. 9.18).
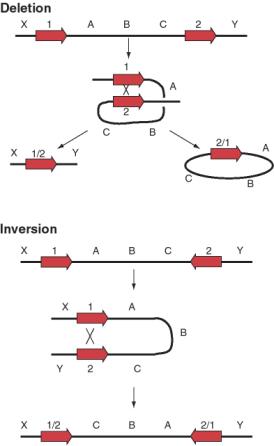
Figure 9.18. Possible
outcomes of recombination between two transposable elements.
The preference of the retrotransposition machinery
for LINE1 mRNA does not appear to be absolute. Many processed genes have been found in eukaryotic
genomes; these are genes that have no introns. In many cases, a homologous gene
with introns is seen in the genome, so it appears that these processed genes
have lost their introns. It is likely that these were formed when processed
mRNA derived from the homologous gene with introns was copied into cDNA and
reinserted into the genome. Many, but not all, of these processed genes are
pseudogenes, i.e. they have been mutated such that they no longer encode
proteins. Other examples of active processed genes have inserted next to
promoters and encode functional proteins.
Additional Readings
Shapiro, J. A. (editor) (1983) Mobile Genetic Elements (Academic Press, Inc., New York).
Fedoroff, N. and Botstein, D. (1992) The Dynamic Genome: Barbara McClintock’s Ideas in the Century of Genetics (Cold Spring Harbor Press, Plainview, NY).
McClintock, B. (1952) Chromosome organization and genic expression. Cold Spring Harbor Symposium on Quantitative Biology 16: 13-47.
Fedoroff, N., Wessler, S., and Shure, M. (1983) Isolation of the transposable maize controlling elements Ac and Ds. Cell 35:235-242.
Boeke, J.D., Garfinkel, D.J., Styles, C.A., and Fink G.R. (1985) Ty elements transpose through an RNA intermediate. Cell 40:491-500
Kazazian, H.H. Jr, Wong, C., Youssoufian, H., Scott, A.F., Phillips, D.G., and Antonarakis, S.E. (1988) Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 332:164-166.
International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409: 860-921. The material from pages 879-889 covers human repeats and transposable elements.
CHAPTER
9
TRANSPOSITION
Questions
Question 9.5. Suppose you are studying a gene that is contained within a 5 kb EcoRI fragment for the wild type allele. When analyzing mutations in that gene, you found one that converted the 5 kb fragment to an 8 kb EcoRI fragment. Further analysis showed that the additional 3 kb of DNA was flanked by direct repeats of 6 bp, that the terminal 30 bp of the additional DNA was identical at each end but in an inverted orientation. Recombinant plasmids carrying the 8 kb EcoRI fragment conferred resistance to the antibiotic kanamycin in the host bacteria, whereas neither the parental cloning vector nor a recombinant plasmid carrying the 5 kb EcoRI fragment did. What do you conclude is the basis for this mutation? What other enzyme activities might you expect to be encoded in the additional DNA?
Use the following diagram to answer the next two questions. Transposase encoded by a transposable element (TE) has nicked on each side of the TE in the donor (black) replicon and made a staggered break in the recipient (gray) replicon, and the ends of the TE have been joined to the target (T) site in the recipient replicon. The strands of the replicons have been designated top (t) or bottom (b). The open triangles with 1 or 2 in them just refer to locations in the figure; they are not part of the structure.
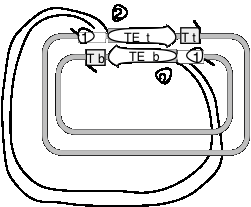
Question 9.6. The action of DNA polymerase plus dNTPs, primed at positions 1, followed by ligase (with ATP or NAD) leads to what product or result? (In this scenario, nothing occurs at positions 2).
Question 9.7. The action of an endonuclease at the positions labeled 2 followed by DNA polymerase and dNTPs to fill in the gaps (from positions 1 to the next 5' ends of DNA fragments), and finally DNA ligase (with ATP or NAD) leads to what product or result?
Question 9.8. Refer to the model for a crossover intermediate in replicative transposition in Fig. 9.13. If the transposon moved to a second site on the same DNA molecule by replicative transposition (not to a different molecule as shown in the Figure), what are the consequences for the DNA between the donor and recipient sites?
Question 9.9. The technique of transposon tagging uses the integration of transposons to mutate a large numbers of genes while leaving a "tag" in the mutated gene to allow subsequent isolation of the gene using molecular probes (such as hybridization probes for the transposon). What is a good candidate for transposon tagging in mammalian cells?