CHAPTER 3
ISOLATING AND
ANALYZING GENES
Recombinant DNA,
Polymerase Chain Reaction and Applications to Eukaryotic Gene Structure and
Function
The first two chapters covered many important aspects of genes, such as how they function in inheritance, how they code for protein (in general terms) and their chemical nature. All this was learned without having a single gene purified. A full understanding of a gene, or the entire set of genes in a genome, requires that they be isolated and then studied intensively. Once a gene is “in hand”, in principal one can determine both its biochemical structures and its function(s) in an organism. One of the goals of biochemistry and molecular genetics is to assign particular functions to individual or composite structures. This chapter covers some of the techniques commonly used to isolate genes and illustrates some of the analyses that can be done on isolated genes.
Methods to purify some abundant proteins were developed early in the 20th century, and some of the experiments on the fine structure of the gene (colinearity of gene and protein for trpA and tryptophan synthase) used microbial genetics and proteins sequencing. However, methods to isolate genes were not developed until the 1960’s, and the were applicable to only a few genes.
All this changed in the late 1970’s with the development of recombinant DNA technology, or molecular cloning. This technique enabled researchers to isolate any gene from any organism from which one could isolate intact DNA (or RNA). The full potential to provide access to all genes of organisms is now being realized as full genomes are sequenced. One of the by-products of the intense investigation of individual DNA molecules after the advent of recombinant DNA was a procedure to isolate any DNA for which one knows the sequence. This technique, called the polymerase chain reaction (PCR), is far easier than traditional molecular cloning methods, and it has become a staple of many laboratories in the life sciences. After covering the basic techniques in recombinant DNA technology and PCR, their application to studies of eukaryotic gene structure and function will be discussed.
Like many advances in molecular genetics, recombinant DNA technology has its roots in bacterial genetics.
Transducing phage
The first genes isolated were bacterial genes that could be picked up by bacteriophage. By isolating these hybrid bacteriophage, the DNA for the bacterial gene could be recovered in a highly enriched form. This is the basic principal behind recombinant DNA technology.
Some bacteriophage will integrate into a bacterial chromosome and reside in a dormant state (Fig. 3.1). The integrated phage DNA is called a prophage, and the bacterium is now a lysogen. Phage that do this are lysogenic. Induction of the lysogen will result in excision of the prophage and multiplication to produce many progeny, i.e. it enters a lytic phase in which the bacteria are broken open and destroyed. The nomenclature is descriptive. The bacteria carrying the prophage show no obvious signs of the phage (except immunity to superinfection with the same phage, covered later in Part Four), but when induced (e.g. by stress or UV radiation) they will generate a lytic state, hence they are called lysogens. Induced lysogens make phage from the prophage that was integrated. Phage that always multiply when they infect a cell are called lytic.
Excision of a prophage from a lysogen is not always precise. Usually only the phage DNA is cut out of the bacterial chromosome, but occassionally some adjacent host DNA is included with the excised phage DNA and encapsidated in the progeny. These transducing phage are usually biologically inactive because the piece of the bacterial chromosome replaces part of the phage chromosome; these can be propagated in the presence of helper phage that provide the missing genes when co-infected into the same bacteria. When DNA from the transducing phage is inserted into the newly infected cell, the bacterial genes can recombine into the host chromosome, thereby bringing in new alleles or even new genes and genetically altering the infected cell. This process is called transduction.
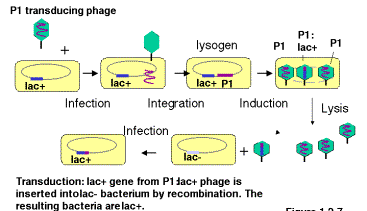
Figure 3.1. Transfer of bacterial genes by transduction: A lac+ transducing phage can convert a lac‑ strain to lac+ by infection (and subsequent crossing over).
Note that the transducing phage are carrying one or a small number of bacterial genes. This is a way of isolating the genes. The bacterial gene in the transducing phage has been separated from the other 4000 bacterial genes (in E. coli). By isolating large numbers of the transducing phage, the phage DNA, including the bacterial genes, can be obtained in large quantities for biochemical investigation. One can isolate mg or mg quantities of a single DNA molecule, which allows for precise structural determination and detailed investigation.
A generalized
transducing phage can integrate at many
different locations on the bacterial chromosome. Imprecise excision from any of
those locations generates a particular transducing phage, carrying a short
sections of the bacterial genome adjacent to the integration site. Thus a
generalized transducing phage such as P1 can pick up many different parts of
the E. coli genome.
A specialized transducing phage integrates into only one or very few sites in the
host genome. Hence it can carryonly a few specific bacterial genes, e.g., l lac
(Fig. 3.2).

Figure 3.2. An example of a l transducing phage carrying part of the lac operon.
This process of isolating a particular bacterial gene on a transducing phage is mimicked in recombinant DNA technology, in which a gene or genome fragment from any organism is isolated on a recombinant phage or plasmid.
Overview of Recombinant DNA Technology
Recombinant DNA technology utilizes the power of microbiological selection and screening procedures to allow investigators to isolate a gene that represents as little as 1 part in a million of the genetic material in an organism. The DNA from the organism of interest is divided into small pieces that are then placed into individual cells (usually bacterial). These can then be separated as individual colonies on plates, and they can be screened through rapidly to find the gene of interest. This process is called molecular cloning.
Joining DNA in vitro to form recombinant molecules
Restriction endonucleases cut at defined sequences of (usually) 4 or 6 bp. This allows the DNA of interest to be cut at specific locations. The physiological function of restriction endonucleases is to serve as part of system to protect bacteria from invasion by viruses or other organisms. (See Chapter 7)
Table 3.1. List of restriction endonucleases and their cleavage sites.
A ' means that the nuclease cuts between these 2 nucleotides to generate a 3' hydroxyl and a 5' phosphate.
|
Enzyme |
Site |
|
Enzyme |
Site |
|
AluI |
AG'CT |
|
NotI |
GC'GGCCGC |
|
BamHI |
G'GATCC |
|
PstI |
CTGCA'G |
|
BglII |
A'GATCT |
|
PvuII |
CAG'CTG |
|
EcoRI |
G'AATTC |
|
SalI |
G'TCGAC |
|
HaeIII |
GG'CC |
|
Sau3AI |
'GATC |
|
HhaI |
GCG'C |
|
SmaI |
CCC'GGG |
|
HincII |
GTY'RAC |
|
SpeI |
A'CTAGT |
|
HindIII |
A'AGCTT |
|
TaqI |
T'CGA |
|
HinfI |
G'ANTC |
|
XbaI |
T'CTAGA |
|
HpaII |
C'CGG |
|
XhoI |
C'TCGAG |
|
KpnI |
GGTAC'C |
|
XmaI |
C'CCGGG |
|
MboI |
'GATC |
|
|
|
N = A,G,C or T
R = A or G
Y = C or T
S = G or C
W = A or T
a. Sticky
ends
(1) Since the recognition sequences for restriction endonucleases are pseudopalindromes, an off-center cleavage in the recognition site will generate either a 5' overhang or a 3' overhang with self-complementary (or "sticky") ends.
e.g. 5' overhang EcoRI G'AATTC
BamHI G'GATCC
3' overhang PstI CTGCA'G
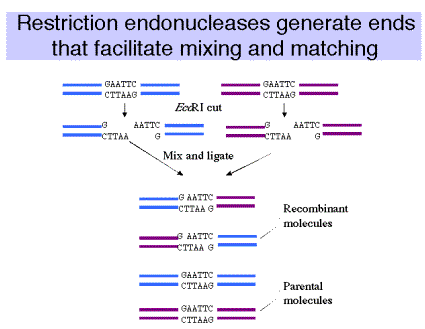
(2) When the ends of the restriction fragments are complementary,
e.g. for EcoRI 5'‑‑‑G AATTC‑‑‑3'
3'‑‑‑CTTAA G‑‑‑5'
the ends can anneal to each other. Any two fragments, regardless of their origin (animal, plant, fungal, bacterial) can be joined in vitro to form recombinant molecules (Fig. 3.3).
Figure 3.3.
b. Blunt
ends
(1) The restriction endonuclease cleaves in the center of the pseudopalindromic recognition site to generate blunt (or flush) ends.
(2) E.g. HaeIII GG'CC
HincII GTY'RAC
T4 DNA ligase is used to tie together fragments of DNA (Fig. 3.4). Note that the annealed "sticky" ends of restriction fragments have nicks (usually 4 bp apart). Nicks are breaks in the phosphodiester backbone, but all nucleotides are present. Gaps in one strand are missing a string of nucleotides.
T4 DNA ligase uses ATP as source of adenylyl group attached to 5' end of the nick, which is a good leaving group after attack by the 3' OH. (See Chapter 5 on Replication).
At high concentration of DNA ends and of ligase, the enzyme can also ligate together blunt‑ended DNA fragments. Thus any two blunt‑ended fragments can be ligated together.
Note: Any fragment with a 5' overhang can be readily converted to a blunt‑ended molecule by fill‑in synthesis catalyzed by a DNA polymerase (often the Klenow fragment of DNA polymerase I). Then it can be ligated to another blunt‑ended fragment.

Figure 3.4
Linkers are short duplex oligonucleotides that contain a restriction endonuclease cleavage site. They can be ligated onto any blunt‑ended molecule, thereby generating a new restriction cleavage site on the ends of the molecule. Ligation of a linker on a restriction fragment followed by cleavage with the restriction endonuclease is one of several ways to generate an end that is easy to ligate to another DNA fragment.
Annealing of homopolymer tails are another way to joint two different DNA molecules.
The enzyme terminal deoxynucleotidyl transferase will catalyze the addition of a string of nucleotides to the 3' end of a DNA fragment. Thus by incubating each DNA fragment with the appropriate dNTP and terminal deoxynucleotidyl transferase, one can add complementary homopolymers to the ends of the DNAs that one wants to combine. E.g., one can add a string of G's to the 3' ends of one fragment and a string of C's to the 3' ends of the other fragment. Now the two fragments will join together via the homopolymer tails.
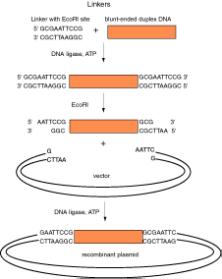
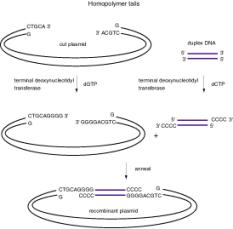
Figure 3.5. Use of linkers (left) and homopolymer tails (right) to make recombinant DNA molecules.
Introduction of recombinant DNA into cell and replication: Vectors
Vectors used to move DNA between
species, or from the lab bench into a living cell, must meet three requirements
(Fig. 3.6).
(1) They must be autonomously replicating DNA molecules in the host cell. The most common vectors are designed for replicating in bacteria or yeast, but there are vectors for plants, animals and other species.
(2) They must contain a selectable marker so cells containing the recombinant DNA can be distinguished from those that do not. An example is drug resistance in bacteria.
(3) They must have an insertion site to accomodate foreign DNA. Usually a unique restriction cleavage site in a nonessential region of the vector DNA. Later generation vectors have a set of about 15 or more unique restriction cleavage sites.
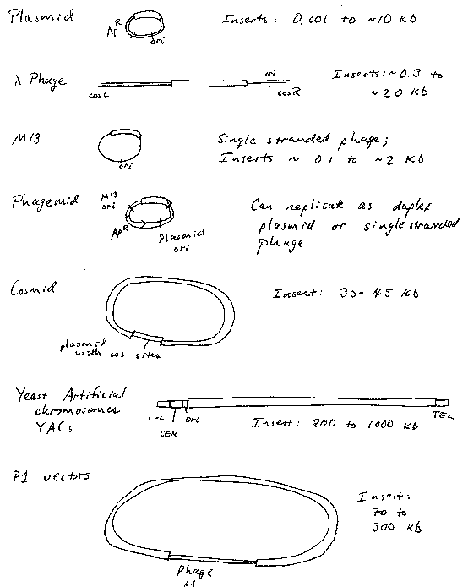
Figure 3.6. Summary of vectors for molecular cloning
Plasmid vectors
Plasmids are autonomously replicating circular DNA molecules found in bacteria. They have their own origin of replication, and they replicate independently of the origins on the "host" chromosome. Replication is usually dependent on host functions, such as DNA polymerases, but regulation of plasmid replication is distinct from that of the host chromosome. Plamsids, such as the sex-factor F, can be very large (94 kb), but others can be small (2‑4 kb). Plasmids do not encode an essential function to the bacterium, which distinguishes them from chromosomes.
Plasmids can be present in a single copy, such as F, or in multiple copies, like those used as most cloning vectors, such as pBR322, pUC, and pBluescript.
In nature, plasmids provide carry some useful function, such as transfer (F), or antibiotic resistance. This is what keeps the plasmids in a population. In the absence of selection, plasmids are lost from bacteria.
The antibiotic resistance genes on plasmids are often carried within, or are derived from, transposons, a types of transposable element. These are DNA segments that are capable of "jumping" or moving to new locations (see Chapter 9).
A plasmid that was widely used in many recombinant DNA projects is pBR322 (Fig. 3.7). It replicates from an origin derived from a colicin-resistance plasmid (ColE1). This origin allows a fairly high copy number, about 100 copies of the plasmid per cell. Plasmid pBR322 carries two antibiotic resistance genes, each derived from different transposons. These transposons were initially found in R-factors, which are larger plasmids that confer antibiotic resistance.
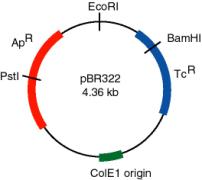
Figure 3.7. Features of plasmid pBR322. The gene conferring resistance to ampicillin (ApR) can be interrupted by insertion of a DNA fragment into the PstI site, and the gene conferring resistance to tetracycline (TcR) can be interrupted by insertion of a DNA fragment into the BamHI site. Replication is controlled by the ColE1 origin.
Use of the TcR and ApR genes allows for easy screening for recombinants carrying inserts of foreign DNA. For instance, insertion of a restriction fragment in the BamHI site of the TcR gene inactivates that gene. One can still select for ApR colonies, and then screen to see which ones have lost TcR .
Question 3.1. What effects on drug resistance are seen when you use the EcoRI or PstI sites in pBR322 for inserting foreign DNA?
A generation of vectors developed
after pBR322 are designed for even more efficient screening for recombinant
plasmids, i.e. those that have foreign DNA
inserted. The pUC plasmids
(named for plasmid universal cloning) and plasmids derived
from them use a rapid screen for inactivation of the b-galactosidase gene to identify recombinants
(Fig. 3.8).
One can screen for production of functional b‑galactosidase in a cell by using the chromogenic substrate X‑gal (a halogenated indoyl b‑galactoside). When cleaved by b‑galactosidase, the halogenated indoyl compound is liberated and forms a blue precipitate. The pUC vector has the b‑galactosidase gene {actually only part of it, but enough to form a functional enzyme with the rest of the gene that is encoded either on the E. coli chromosome or an F' factor}. When introduced into E. coli, the colonies are blue on plates containing X‑gal.
The multiple cloning sites (unique restriction sites) are in the b‑galactosidase gene (lacZ). When a restriction fragment is introduced into one or more of these sites, the b‑galactosidase activity is lost by this insertional mutation. Thus cells containing recombinant plasmids form white (not blue) colonies on plates containing X‑gal.
The replication origin is a modified ColE1 origin of replication that has been mutated to eliminate a negative control region. Hence the copy number is very high (several hundred or more plasmid molecules per cell), and one obtains an very high yield of plasmid DNA from cultures of transformed bacteria. The plasmid has ApR as a selectable marker.
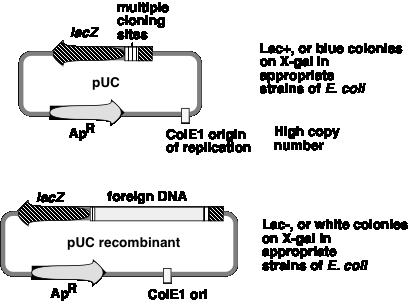
Figure 3.8. pUC-type vectors
Introduction of a recombinant DNA molecule into a host cell
Introduction into CaCl2 treated E. coli: transformation
E. coli does not have a natural system for taking up DNA, but when treated with CaCl2, the cells will take up the added DNA (Fig. 3.9). The recombinant vectors will give a new phenotype to the cells (usually drug resistance), so this process can be considered DNA-mediated transformation. An average efficiency is about 106 transformants per mg of DNA, although some more elaborate transformation cocktails procedures can give up to about 108 transformants per mg of DNA.
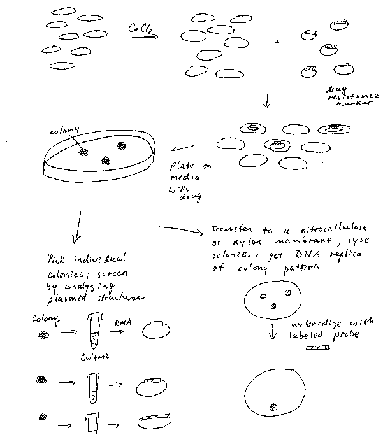
Figure 3.9. DNA-mediated
transformation of E. coli.
Usually one will transform with a mixture of recombinant vector molecules, most of which carry a different restriction fragment. Each transformed E. coli cell will pick up only one plasmid molecule, so the complex mixture of plasmids in the ligation mix has been separated into a population of transformed bacteria (Fig. 3.9). The bacterial cells are then plated at a sufficiently low density that individual colonies can be identified. Each colony (or transformant) carries a single plasmid, so as one screens the colonies, one is actually screening through individual DNA molecules. A colony is a visible group of bacterial cells on a plate, all of which are derived from a single bacterial cell. A group of identical cells derived from a single cell is called a clone. Since each clone carries a single type of recombinant DNA molecule, the process is called molecular cloning.
Phage vectors for more
efficient introduction of DNA into bacteria.
Phage vectors such as those derived from bacteriophage l can carry larger inserts and can be introduced into bacteria more efficiently. l phage has a duplex DNA genome of about 50 kb. The internal 20 kb can be replaced with foreign DNA and still retain the lytic functions. Hence restriction fragments up to 20 kb can replace the l sequences, allowing larger genomic DNA fragments to be cloned (Fig. 3.10).
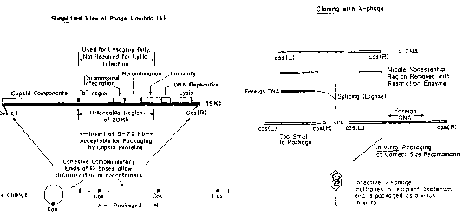
Figure 3.10. Lambda vectors for cloning.
Recombinant bacteriophage can be introduction into E. coli by infection. DNA that has the cohesive ends of l can be packaged in vitro into infective phage particles. Being in a viral particle brings the efficiency of infection reliably over 108 plaque forming units per mg of recombinant DNA.
Some other bacteriphage vectors for cloning are derived from the virus M13. One can obtain single stranded DNA from M13 vectors and recombinants. M13 is a virus with a genome of single stranded DNA. It has a nonessential region into which foreign genes can be inserted. It has been modified to carry a gene for b‑galactosidase as a way to screen for recombinants. Introduction of recombinant M13 DNA into E. coli will lead to an infection of the host, and the progeny viral particles will contain single‑stranded DNA. The replicative form is duplex, allowing one to cleave with restriction enzymes and insert foreign DNA.
Some vectors are hybrids between plasmids and single‑strand phage; these are called phagemids. One example is pBluescript. Phagemids are plasmids (with the modified, high-copy number ColE1 origin) that also have an M13 origin of replication. Infection of transformed bacteria (containing the phagemid) with a helper virus (e.g. derived from M13) will cause the M13 origin to be activated, and progeny viruses carrying single‑stranded copies of the phagemid can be obtained. Hence one can easily obtain either double‑ or single‑stranded forms of thes plasmids. {The "blue" comes from the blue‑white screening for recombinants that can be done when the multiple cloning sites are in the b‑galactosidase gene. The "script" refers to the ability to make RNA copies of either strand in vitro with phage RNA polymerases.}
Vectors designed to carry larger inserts
Fragments even larger than those carried in l vectors are useful for studies of longer segments of chromosomes or whole genomes. Several vectors have been designed for cloning these very large fragments, 50 to 400 kb.
Cosmids are plasmids that have the cohesive ends of l phage. They can be packaged in vitro into infective phage particles to give a more efficient delivery of the DNA into the cells. They can carry about 35 to 45 kb inserts (Fig. 3.6).
Yeast artificial chromosomes (YACs) are yeast vectors with centromeres and telomeres. They can carry about 200 kb or larger fragments (in principle up to 1000 kb = 1 Mb). Thus very large fragments of DNA can be cloned in yeast (Fig. 3.11). In practice, chimeric clones with fragments from different regions of the genome are obtained fairly often, and some of the inserts are unstable.
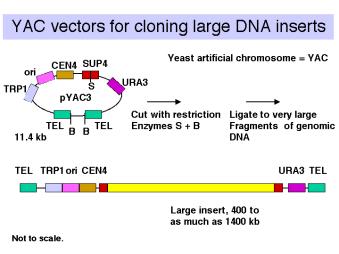
Figure 3.11
Vectors derived from bacteriophage P1 can carry fragments of about 100 kb. Fragments in a similar size range are also cloned into bacterial artificial chromosomes (BACs), which are derived from the F-factor (Fig. 3.12). These have a lower copy number (like F) but they are stable and relatively easy to work with in the laboratory. BACs have become one of the most frequently used vectors for large inserts in genome projects.
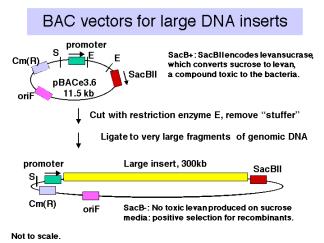
Figure 3.12.
Shuttle vectors for testing functions of isolated genes
Shuttle vectors can replicate in two different organisms, e.g. bacteria and yeast, or mammalian cells and bacteria. They have the appropriate origins of replication. Hence one can, e.g. clone a gene in bacteria, maybe modify it or mutate it in bacteria, and test its function by introducing it into yeast or animal cells.
Polymerase Chain Reaction, or PCR
The polymerase chain reaction, or PCR, is now one of the most commonly used assays for obtaining a particular segment of DNA or RNA. It is rapid and extremely sensitive. By amplifying a designated segment of DNA, it provides a means to isolate that particular DNA segment or gene. This method requires knowledge of the nucleotide sequence at the ends of the region that you wish to amplify. Once that is known, one can make large quantities of that region starting with miniscule amounts of material, such as the DNA within a single human hair. With the availability of almost complete or complete sequences of genomes from many species, the range of genes to which it can be applied is enormous. The applications of PCR are numerous, from diagnostics to forensics to isolation of genes to studies of their expression.
The power of PCR lies in the exponential increase in amount of DNA that results from repeated cycles of DNA synthesis from primers that flank a given region, one primer designed to direct synthesis complementary to the top strand, the other designed to direct synthesis complementary to the bottom strand (Fig. 3.13. When this is done repeatedly, there is roughly a 2-fold increase in the amount of synthesized DNA in each cycle. Thus it is possible to generate a million-fold increase in the amount of DNA from the amplified region with a sufficient number of cycles. This exponential increase in abundance is similar to a chemical chain reaction, hence it is called the polymerase chain reaction.

Figure 3.13. Polymerase Chain Reaction (PCR)
The events in the polymerase chain reaction are examined in more detail in Fig. 3.14. The several panels show what happens in each cycle. Each cycle consists of a denaturation step at a temperature higher than the melting temperature of the duplex DNA (e.g. 95oC ), then an annealing step at a temperature below the melting temperature for the primer-template (e.g. 55oC), followed by extension of the primer by DNA polymerase using dNTPs provided in the reaction. This is done at the temperature optimum for the DNA polymerase (e.g. 70oC for a thermostable polymerase). Thermocylers are commercially available for carrying out many cycles quickly and reliably.
The template supplied for the reaction is the only one availablein the first cycle, and it is still a major template in the second cycle. At the end of the second cycle, a product is made whose ends are defined by primers. This is the desired product, and it serves as the major template for the remaining cycles. The initial template is still present and can be used, but it does not undergo the exponential expansion observed for the desire product.
If n is the number of cycles, the amount of desired product is approximately 2n-1 –2 times the amount of input DNA (between the primers). Thus in 21 cycles, one can achieve a million-fold increase in the amount of that DNA (assuming all cycles are completely efficient). A sample with 0.1 pg of the segment of DNA between the primers can be amplified to 0.1 mg in 21 cycles, in theory. In practice, roughly 25-35 cycles are done in many PCR assays.
The ease if doing PCR was greatly increased by the discovery of DNA polymerases that were stable at high temperatures. These have been isolated from bacteria that grow in hot springs, such as those found in Yellowstone National Park, such as Thermus aquaticus. The Taq polymerase from this bacterium will retain activity even at the high temperatures needed for melting the templates, and it is active at a temperature between the melting and annealing temperature. This particular polymerase is rather error-prone, and other thermostable polymerases have been discovered that are more accurate.
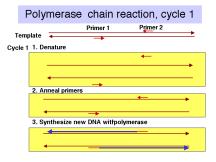
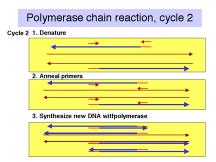
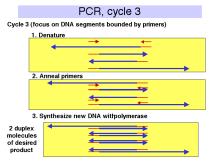
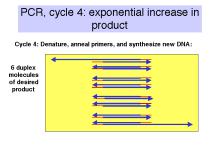
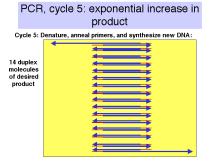
Figure 3.14. Steps
in the polymerase chain reaction.
cDNA clones are copies of mRNAs
Construction of cDNA clones involves the synthesis of complementary DNA from mRNA and then inserting a duplex copy of that into a cloning vector, followed by transformation of bacteria (Fig. 3.15).
a. First strand synthesis:
First, one anneals an oligo dT primer onto the 3' polyA tail of a population of mRNAs. Then reverse transcriptase will begin DNA synthesis at the primer, using dNTPs supplied in the reaction, and copy the mRNA into complementary DNA, abbreviated cDNA.
The mRNA is degraded by the RNase H activity associated with reverse transcriptase and by subsequent treatment with alkali.
b. Second strand synthesis:
For the primer to make the second strand of DNA (equivalent in sequence to the original mRNA), one can utilize a transient hairpin at the end of the cDNA. (The basis for its formation is not certain.) In other schemes, one generates a primer binding site and uses a primer directed to that site; one way to do this is by homopolymer tailing of the cDNA followed by use of a complementary primer. Random primers can also be used for second strand synthesis; although this precludes the generation of a full-length cDNA (i.e. a copy of the entire mRNA). However, it is rare to generate duplex copies of the entire mRNA by any means.
DNA polymerase (e.g. Klenow polymerase) is used to synthesize the second strand, complementary to the cDNA. The product is duplex cDNA.
If the hairpin was used to prime second strand synthesis, it must be opened by a single‑strand specific nuclease such as S1.
c. Insertion of the duplex cDNA into a cloning vector:
One method is to use terminal deoxynucleotidyl transferase to add a homopolymer such as poly-dC to the ends of the duplex cDNA and a complementary homopolymer such as poly-dG to the vector.
An alternative approach is to use linkers; these can be employed such that a linker carrying a cleavage site for one restriction endonuclease is on the 5' end of the duplex cDNA and a linker carrying a cleavage site for a different restriction endonuclease is on the 3' end. (In this context, 5’ and 3’ refer to the nontemplate, or "top" strand.) This allows "forced" cloning into the vector, and one has initial information about orientation, based on proximity to one cleavage site or the other.
The cDNA and vector are joined at the ends, using DNA ligase, to form recombinant cDNA plasmids (or phage).
d. The ligated cDNA plasmids are then transformed into E. coli. The resulting set of transformants is a library of cDNA clones.
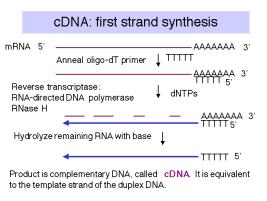
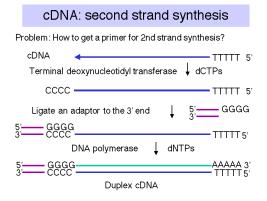
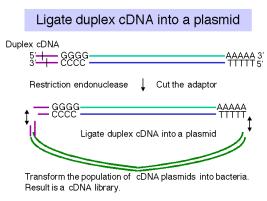
Figure 3.15. Making cDNA clones
Screening methods for cDNA clones
a. Brute
force examination of individual cDNA plasmids.
If the mRNA is highly abundant in a given tissue, then many of the cDNA clones will be copies of that mRNA. One can examine DNA from individual clones and test for characteristic restriction cleavage patterns or a particular sequence. This was a common approach for screening cDNAs in the early days of recombinant DNA technology.
Starting in the mid-1990’s, cooperative efforts from corporations (such as Merck) and publicly funded genome centers (such as at Washington University) have generated the sequence of individual clones from large cDNA libraries from many tissues from human, mouse, and rat. Other consortia have sequenced cDNA libraries from other species. Each sequence is called an “expressed sequence tag” or EST. These are now a major source of partially or fully characterized cDNA clones. Hundreds of thousands of ESTs are available, and contain at part of the DNA sequence from many, if not most, human genes. The web site for NCBI (http://www.ncbi.nlm.nih.gov) is an excellent resource for examining the ESTs.
b. Hybridization
with a gene‑specific probe.
If the sequence of the desired cDNA is known, or if the sequence from homologs from related species is known, one can use synthetic oligonucleotides (or other source of the diagnostic sequence) as a radiolabeled hybridization probe to identify the cDNA of interest.
If the amino acid sequence has been determined for all or even just parts of the protein product of the gene of interest, then one can chemically synthesize oligonucleotides based on the genetic code for those amino acids. The oligonucleotides need to be at least 18 nucleotides or longer (so that they will anneal to specific sites in the genome), and because the genetic code is degenerate (more than one codon per amino acid; discussed in Part Two), they have to be degenerate as well. The oligonucleotides can be used directly as hybridization probes, although it is becoming more common to amplify the region between two oligonucleotides using the polymerase chain reaction, and to use that amplification product as a labeled probe.
The process of hybridization screening is illustrated schematically in Fig. 3.16. The colonies of bacteria, each with a single cDNA plasmid, are transferred to a solid substrate (such as a nylon or nitrocellulose membrane), lysed. and the released DNA immobilized onto the membrane. Hybridization of this membrane (with the DNA attached) to a specfic probe allows one to screen through thousands of colonies in a single experiment.
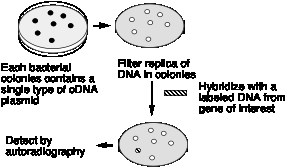
Figure 3.16 Hybridization Screening
c. Express the cDNA, i.e. make the protein product encoded by the mRNA, and screen for that protein product (Fig. 3.17). This is often in bacteria by constructing the clones in a vector that has an active E. coli promoter (for transcription) and efficient translation signals upstream from the site at which the cDNAs were inserted. The transformed bacterial cells will express the encoded protein, and one tries to identify it. One can also screen for expression in yeast, plant or mammalian cells. The expression vector has to contain gene-regulatory signals (such as promoters and enhancers, see Part Three) that allow expression of the desired gene in the appropriate cell.
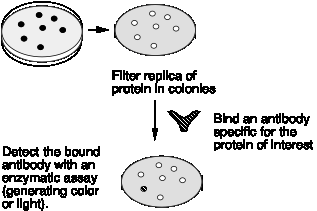
Figure 3.17. Screening for an Expressed Gene Product
(1) One can use specific antisera to detect the desired colony expressing the gene of interest.
(2) One can use a labeled ligand that will bind to the expressed cDNA on the cell surface. For example, cDNAs for receptors can be expressed in an appropriate cell (usualy mammalian cells in culture) and identified by newly-acquired ability to bind a labeled hormone (such as growth hormone or erythropoietin)
(3) by complementation of a known mutation in the host. E.g. a cDNA for the human homolog to yeast p34cdc2 was isolated by its ability to complement a yeast mutant that had lost the function of this key regulator of progress through the cell cycle.
(4) Expression cloning can be done in mammalian cells, as long as one can screen or select for a new function generated by the expression. Use of this method to isolate the receptor for the glycoprotein hormone erythropoietin is illustrated in Fig. 3.18.
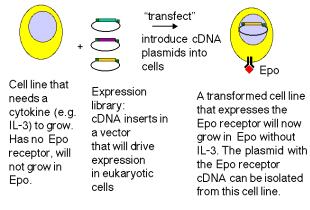
Figure 3.18. Expression screening in eukaryotic cells.
d. Differential analysis:
Often one is interested in finding all the genes (or their mRNAs) that are expressed uniquely in some differentiated or induced state of cells. Two classic examples are (i) identifying the genes whose products regulate the determination process that causes a multipotential mouse cell line (like 10T1/2 cells) to differentiate into muscle cells, and (ii) ,using the fact that the T-cell receptor is expressed only in T-lymphocytes, but not in their sister lineage B-lymphocytes, to help isolate cDNA clones for that mRNA. Both of these projects used subtractive hybridization to highly enrich for the cDNA clones of interest.
In this technique, the cDNA from the differentiating or induced cell of interest is hybridized to mRNA from a related cell line, but which has not undergone the key differentiation step. This allows one to remove mRNA-cDNA duplexes that contain the cDNAs for all the genes expressed in common between the two types of cells. The resulting single-stranded are enriched for the cDNAs that are involved in the process under study.
The subtractive hybridization scheme used in isolation of the muscle determination gene MyoD is illustrated in Fig. 3.19.
A conceptually equivalent strategy, using PCR (see next section) rather than cDNA cloning, is differential display of PCR products from cells that differ by some process (e.g. differentiation, induction, growth arrest versus stimulation, etc.). In this technique, one uses several sets of PCR primers annealed to cDNA to mRNA from the two types of cells that are being compared. The sets of primers are empirically designed to allow many regions of cDNA to be amplified. The amplification products are resolved (or displayed) on polyacrylamide gels, and the products specific to the cell type of interest are isolated and used to screen through cDNA libraries. This technique is also called representational difference analysis.
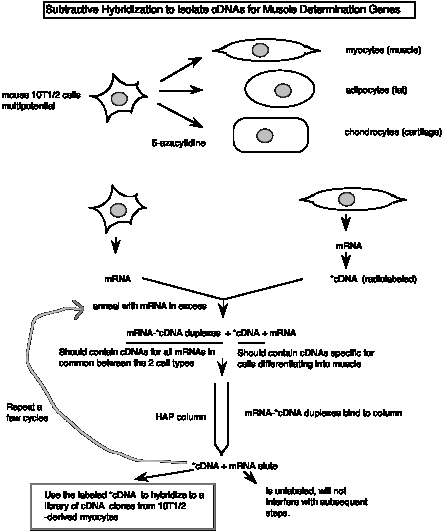
Figure 3.19. Differential screening to find cDNAs of mRNAs expressed only in certain cell-types.
The advent of sequencing all or a very large number of genes from various organisms (e.g. E. coli, yeast, Drosophila, humans) has allowed the development of high-density microchip arrays of DNA from each gene. One can hybridize RNA from cells or tissues of interest, isolated under various metabolic conditions, to identify all (known) genes expressed. Even more useful are assays for genes whose expression changes during a shift in cell metabolism (cell cycle, heat shock, hormonal induction, etc.) or as a result of mutation of some other gene (e.g. a gene encoding a transcription factor of interest). This powerful new technology is being used more and more to examine global effects on gene expression.
For a description (and movie) of the Affymetrix GeneChip, go to
http://www.affymetrix.com/technology/index.html
Genomic DNA clones
Clones of genomic DNA, containing individual fragments of chromosomal DNA, are needed for many purposes. Some examples include:
§ to obtain detailed structures of genes,
§ to identify regulatory regions, i.e. DNA sequences needed for correct expression of the gene,
§ to map and analyze alterations to the genome, e.g. the isolate genes that when mutated cause a hereditary disease,
§ to direct alterations in the genome, e.g. by homologous recombination to replace a wild-type allele with a mutant one (to test function of the gene in mouse) or vice versa (to cure a hereditary disease, perhaps eventually in humans).
Construction of libraries of genomic DNA fragments in cloning vectors
Genomic DNA is digested with restriction enzymes (Fig. 3.20.) The more frequently an enzyme cuts (the shorter the recognition sequence), the smaller the average size of DNA fragments. Some enzymes cut very infrequently, such as NotI (8 bp recognition sequence) and can be used to generate very large fragments. Alternatively, one can do a partial digest (not all sites are cleaved) with a particular enzyme and isolate the products that are in the desired size range (e.g. 20 kb). A particularly clever way to do this is to digest partially with Sau3AI or MboI (both cut at 'GATC) and ligate these fragments into vector cut with BamHI (cuts at G'GATCC) ‑ i.e. they have the same sequence in the overhang (or sticky end). In this process one uses vectors that can accomodate large DNA fragments, such as l phage vectors, cosmids, YACs or P1 vectors.

Figure 3.20. Construction of a library of genomic DNA
Screening methods for genomic DNA clones
One method is to use complementation of a mutation in the host to select or screen for the desired gene. This works just like the situation for cDNA clones described above, and it requires that the cloned fragments be expressed in the host cell.
Far more common is to screen by hybridization with gene‑specific probes (Fig. 3.21). Frequently the cDNA clone is found first, and the genomic clone then isolated by hybridization screening (using the cDNA clone as a probe) against a library of genomic DNA fragments.
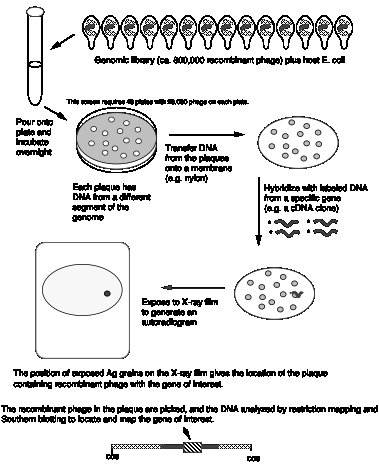
Figure 3.21. Screening a
library of genomic DNA
Eukaryotic gene structure
Much can be learned about any gene after it has been isolated by recombinant DNA techniques. The structure of coding and noncoding regions, the DNA sequence, and more can be deduced. This is true for bacterial and viral genes, as well as eukaryotic cellular genes. The next sections of this chapter will focus on analysis of eukaryotic genes, showing the power of examining purified copies of genes.
Split genes and introns
Precursors to mRNA longer than mRNA
Initial indications of a complex structure to eukaryotic genes came from analysis of nuclear RNAs during the 1970’s. The precursors to messenger RNA, or pre-mRNAs, were found to be surprisingly long, considerably larger than the average mRNA size (Fig. 3.22).
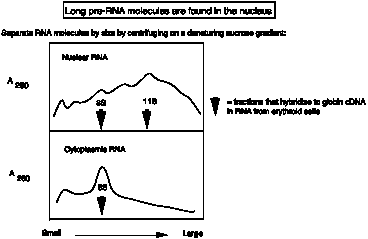
Figure
3.22.
Denaturing sucrose gradients (with high concentration of formamide, e.g. >50%) separate RNAs on the basis of size. Analysis of nuclear RNA showed that the average size was much larger than the average size of cytoplasmic RNA.
Labeled RNA could be "chased" from the nucleus to the cytoplasm ‑ i.e. nuclear RNA was a precursor to mRNA and other cytoplasmic RNAs. Was the extra RNA at the ends? or in the middle of the pre‑mRNA?
More precisely, one could examine specific RNAs by hybridizing fractions from the denaturing sucrose gradients to labeled copies of, e.g. globin mRNA. The hybridizing RNA from the nucleus was about 11S (as well as mature 8S message), whereas cytoplasmic RNA of about 8S hybridized. Thus the nuclear RNA encoding globin is larger than the cytoplasmic mRNA.
Visualization of mRNA-DNA heteroduplexes revealed extra sequences internal to the mRNA-coding segments
R-loops are hybrids between RNA and DNA that can be visualized in the EM, under conditions where DNA‑RNA duplexes are favored over DNA‑DNA duplexes (Fig. 3.23). For a simple gene structure, one sees a continuous RNA‑DNA duplex (smooth, slowly curving) and a displaced single strand of DNA (thinner, many more turns and curves – single stranded DNA is not a rigid as double stranded nucleic acid, either duplex DNA or RNA-DNA).
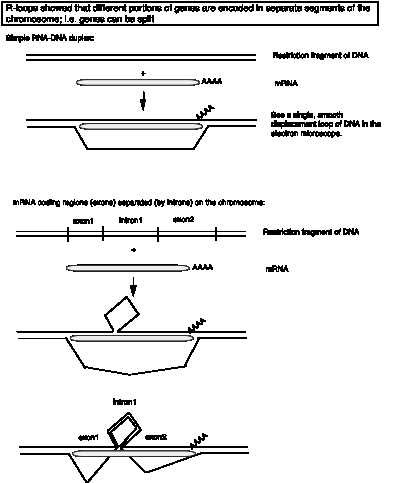

Figure 3.23.
EM pictures of duplexes between purified adenovirus mRNAs and the genomic DNA showed extensions at both the 3' (poly A) and 5' ends, which are encoded elsewhere on the genome. All late mRNAs have the same sequence at the 5' end; this is dervied from from the tripartite leader. R‑loops between late mRNAs and adenovirus DNA fragments including the major late promoter showed duplexes with the leader segments, separated by loops of duplex DNA (Fig. 3.23, bottom panel). The RNA-DNA hybrids identify regions of DNA that encode RNA. The surprising result is that RNA-coding portions of a gene are separated by loops of duplex DNA in the R-loop analysis. Examples of R-loops in genes with introns are shown in Fig. 3.24.
These data showed that the adenovirus RNAs are encoded in different segments of the viral genome; i.e. the genes are split. The portion of a gene that encodes mRNA was termed an exon. The part of gene does not code for sequences in the mature mRNA is called an intron. These observations led to the Nobel Prize for Phil Sharp and Rich Roberts. Louise Chow and Sue Berget were also key players in the discovery of introns.
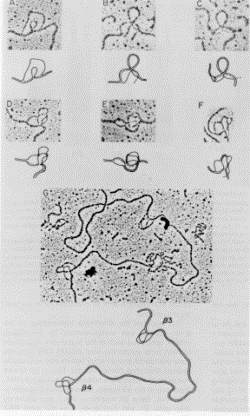
Figure 3.24. R-loops between clones of rabbit beta-like globin
genes (now called HBE and HBG) and mRNA from rabbit embryonic erythroid cells. A
photograph from the electron microscope is shown at the the top of each panel,
and an interpretive drawing is included below it. The displaced nontemplate
strand of DNA forms partial or complete duplexes with the template strand in
the large intron. A small intron is also visible in panel C. Panel G shows the
two genes together on one large clone.
Interruptions in cellular genes were discovered subsequently, in the late 1970's, in globin genes, immunoglobulin genes and others. We now realize that mostgenes in complex eukaryotes are split by multiple introns.
Exons are more conserved than introns (in most cases), since alterations in protein-coding regions that alter or decrease function are selected against, whereas many sequences in introns can be altered without affecting the function of the gene product. Important sequences in introns (such as splice junctions, the branch point, and occassionally enhancers) are covered in some detail in Part Three.
Differences in restiction maps between cDNA and genomic clones reveal introns
Restriction maps based on copies of the mRNA (cDNA) were different from those in genomic DNA ‑ the genes were cleaved by some restriction endonucleases that the cDNAs were not, and some restriction sites were further apart in the genomic DNA. These observations were explained by the presence of intervening sequences or introns (Fig. 3.25).

Figure 3.25.
The experimental procedures to do this involve making a restriction map of the clones of genomic DNA, and then identifying the regions that encode mRNA by hybridization of labeled cDNA probes to the restriction digests. Cloned genomic DNA digested with appropriate restriction endonucleases, separated by size on an agarose gel, and then transferred onto a nylon or nitrocellulose solid support. This Southern blot (see Chapter 2) is then hybridized with a labeled probe specific to the cDNA (composed only of exons). The pattern of labeled fragments on the resulting autoradiogram shows the fragments that contain exons. Alignment of these with the restriction map of the gene gives an approximation of the position of the exons.
The blot-hybridization approach can be combined with a PCR (polymerase chain reaction) analysis for higher resolution. Primers are synthesized that will anneal to adjacent exons. The difference in size of the PCR amplification product between genomic DNA and cDNA is the size of the intron. The PCR product can be cloned and sequenced for more detailed information, e.g. to precisely define the exon/intron junctions.
Subsequently, the nucleotide sequence of exonic regions and preferably the entire gene is determined. The presence of introns were confirmed and their locations defined precisely in DNA sequences of isolated clones of the genes.
Types of exons
Eukaryotic genes are a combination of introns and exons. However, not all exons do the same thing (Fig. 3.26). In particular, the protein-coding regions or genes are a subset of the sequences in exons. Exons include both the untranslated regions and the protein-coding, translated regions. Introns are the segments of genes that are present in the primary transcript (or precursor RNA) but are removed by splicing in the production of mature RNA. Methods used to detect coding regions will not find all exons.
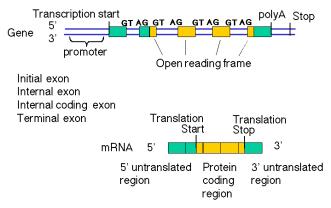
Figure 3.26. Types of exons
Multiple, large introns can make some eukaryotic genes very large
Eukaryotic genes can be split into many (>60), sometimes very small exons (e.g. <60 bp, coding for <20 amino acids), separated by very large introns (as large as >100kb), resulting in some enormous genes (>500 kb). E.g. the DMD gene (which when mutated can cause Duchenne's muscular dystrophy) is almost 1 Mb, about 1/4 the size of the E. coli chromosome!
The average size of genes from more complex organisms is considerably larger than those of simpler ones, but the avg. size of mRNA is about the same, reflecting the presence of more and larger introns in the more complex organisms.
tRNA and rRNA genes also contain introns
Finding exons in long genomic sequences using computer programs
Far more exons and introns have been discovered (or more accurately, predicted) throught the analysis of genomic DNA sequences than could ever be discovered by direct experimentation. The different types of exons, the enormous length of introns, and other factors have complicated the task of finding reliable diagnostic signatures for exons in genomic sequences. However, considerable progress has been made and continues in current research. Some of the commonly used approaches are summarized in Fig. 3.27.


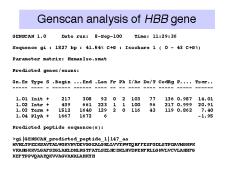
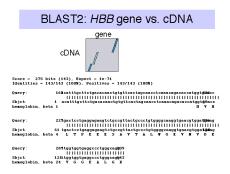
Figure 3.27. Introns in the b-globin
gene can be reliably identified computationally.
Introns are removed by splicing RNA precursors
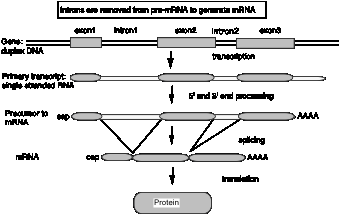
Figure 3.28. Introns are removed from pre-mRNA to generate mRNA.
Alternative splicing generates more than one polypeptide from the same gene
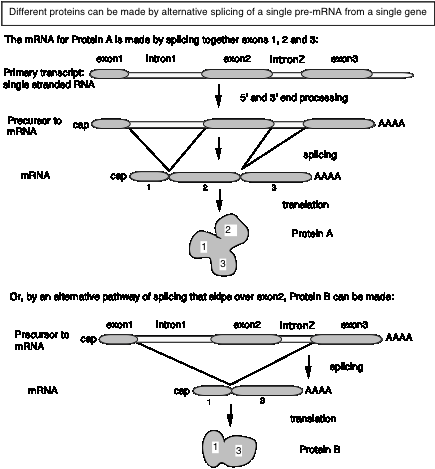
Figure 3.29.
Some segments of RNA may be included in the mature mRNA (exons) but not included on other spliced products. The alternative products may be made in different tissues or at different developmental stages ‑ i.e. alternative splicing can be regulated.
Split genes may enhance the rate of evolution
Many exons encode a unit very close to a protein domain, e.g. the exons of leghemoglobin, or the variable and constant regions of immunoglobulins, or domains (e.g. "kringle") in EGF precursor that are also found in part of the LDL receptor. The exon organization tends to be well conserved in highly divergent species. Introns tend to occur between those portions of genes that encode structural domains of proteins.
Duplication of the exons encoding structural domains and subsequent recombination can lead to more rapid evolution of a new protein, essentially using the parts from earlier evolved genes. Analogous to building a house from prefabricated parts, as opposed to one nail and one board at a time ‑ start with preassembled walls, roof joists etc.
However, the relationship between exons and structural domains of proteins is not exact, and some exon‑intron boundaries vary (a little) in genes for different species. A different model holds that the introns are transposable elements (some certainly are ‑ see later). They can insert anywhere in a gene, but they are least disruptive at domain boundaries, and these latter insertions are more likely to be fixed in a population than insertions into the middle of a region encoding a domain. So the results after long years of evolution is that the introns tend to be between region coding domains, but the gene was originally intact, not assembled from discrete exons.
Multi‑gene families and gene clusters
Many eukaryotic genes are found in multiple copies. Some of them are developmentally regulated, such as HOX gene clusters and globin gene clusters .

Figure
3.30.
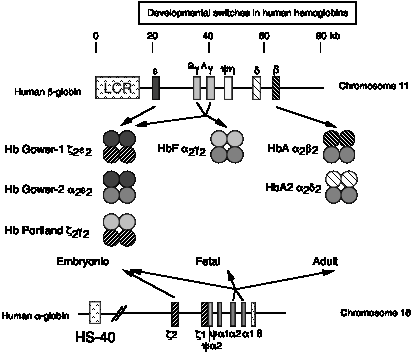
A multigene family contains multiple genes of similar sequence encoding similar proteins; e.g. globin genes (Fig. 3.30). Globin genes are expressed at different times of development. The order of developmental expression is the same as their order along the chromosome, e.g. the e-globin gene is expressed in early embryonic red cells, the g-globin gene is expressed at a high level in fetal red cells, and the b-globin gene is expressed in red cells after birth. As we will see later, this correlates with their distance from a dominant control element at the 5' end of the cluster, the Locus Control Region.
The order of HOX genes is also aligned with their spatial expression in the embryo. This is another example of alignment between chromosomal position and regulation of expression.
Other multi‑gene families include those encoding histones, immunoglobulins, actins, cyclins, cyclin‑dependent protein kinases, and rRNAs. Some of these families are linked in gene clusters, but others are dispersed around the genome. Having multiple copies of genes may be more the rule than the exception in eukaryotic genomes.
Experimental techniques that reveal multigene families include the following.
Purification and analysis of a particular kind of protein, e.g. hemoglobins, immunoglobulins, and many enzymes, may reveal heterogeneity. Further purification (via chromatography and electrophoresis) and sequencing can show that the observed heterogeneity is a result of related but not identical proteins, and one deduces that these similar proteins are encoded by multiple genes with similar sequences, i.e. a multigene family.
Analysis of the clones obtained by screening a library of cloned genomic DNA may reveal multiple related sequences, each with a distinctive restriction map. In many cases these are clones of different, related genes that comprise a multigene family (Fig. 3.31).
Southern blot‑hybridization of restriction‑cleaved genomic DNA can reveal multiple copies of genes, simply as multiple bands on the hybridized blot. Although the number of fragments generated from total genomic DNA is too many to resolve on a gel, after transfer to a membrane, particular fragments can be visualized by hybridization with a specific probe. The number of hybridizing fragments is roughly correlated with the number of copies of related genes. Some genes are cleaved by the restriction enzyme, producing multiple bands, but some fragments can have multiple genes. A true measure of the number of related genes comes from more detailed restriction mapping or sequencing.
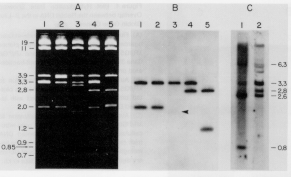
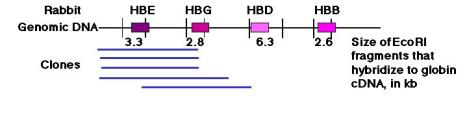
Figure 3.31. Blot-hybridization analysis of clones of genomic DNA and genomic DNA showing that mutliple copies of genes are present. A set of overlapping clones containing rabbit genomic DNA were digested and run on an agarose gel (panel A), blotted onto a membrane and hybridized with a radiolabeled probe that detected embryonic hemoglobin genes, and exposed to X-ray film. The resulting autoradiogram is shownin panel B. Panel C shows the results of a blot-hybridization analysis of rabbit total genomic DNA, using the same probe. Many of the same bands are seen as in the cloned DNA, confirming the existence of multiple hybridizing fragments. Mapping the fragments showed that they represented separate genes.
Keeping multigene families homogeneous
Sometimes multiple copies of genes are maintained as virtually identical over the course of evolution: e.g. rRNA genes, histone genes, a‑globin genes (in primates). In these cases, the multiple copies are coevolving (concerted evolution).
sequence
differences
Human: A | A | A | among human genes: 1%
between human & chimp 5%
Chimp: A | A | A | among chimp genes: 1%
between chimp & monkey 10%
Monkey: A | A | A | among monkey genes: 1%
Since all three primates have 3 A genes, we infer that the common ancestor had 3 genes (the duplications preceded the speciation events). If in the time since human and chimp diverged, the A genes have diverged 5%, why haven't the A genes in human (e.g.) also diverged 5% from each other? They have been apart even longer than the human and chimp chromosomes carrying them! The A genes within a species are "talking to each other", or co‑evolving or evolving in concert.
Sequence homogeneity in a multigene family can arise because of recent gene amplification (Fig. 3.32 part1). In this case the genes have not been separate from each other long enough to accumulate variation in their sequences. Other multigene families have existed for a long time, but maintain sequence homogeneity despite ample opportunity for divergence. Two mechanisms have been seen that maintain similarity. The first is multiple rounds of unequal crossing over. As illustrated in Fig. 3.32, part 2, the expansions and contractions of repeated genes can result in a new variant predominanting in the gene cluster. The other method for maintaining homogeneity is gene conversion between homologs. When a new mutation arises, it can be removed by conversion with the unmutated allele, or the mutation can be passed on the the other allele. Either way, the sequences of the two alleles becomes the same.
Sometimes the products of the gene duplications, or duplicative transpositions, accumulate mutations so they are no longer functional. These remnants of once‑active genes are called pseudogenes.
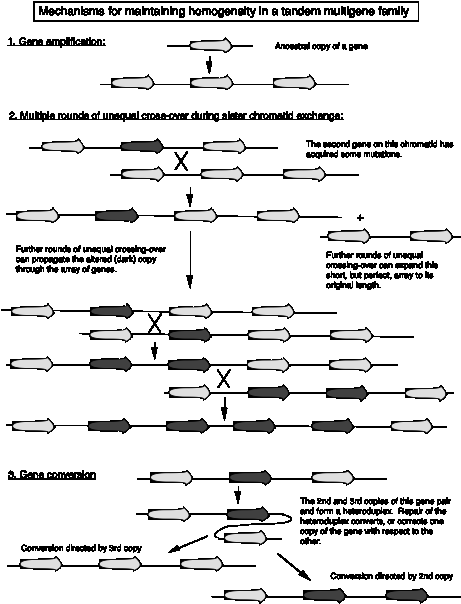

Figure 3.32.
Functional analysis of isolated genes
Gene expression
"Northern blots" or RNA blot‑hybridization
In the reverse of Southern blot‑hybridizations, one can separate RNAs by size on a denaturing agarose gel, and transfer them to nylon or other appropriate solid support. Labeled DNA can then be used to visualize the corresponding mRNA (Fig. 3.33). Ed Southern initially used labeled rRNA to find the complementary regions in immobilized, digested DNA, so this "reverse" of Southern blot-hybridizations, i.e. using a labeled DNA probe to hybridize to immobilized RNA, is often referred to as "Northern" blot‑hybridizations.
One can hybridize a labeled DNA clone to a panel of RNA samples from a wide variety of tissues to determine in what tissues a particular cloned gene is expressed (top panel of Fig. 3.33. More precisely, this technique reveals the tissues in which the genes is transcribed into stable RNA. The results allow one to determine the tissue specificity of expression, e.g. a gene may only be expressed in liver, or only in erythroid cells (e.g. the b-globin gene). This helps give some general idea of the possible function of the gene, since it should reflect the function of that tissue. Other genes are expressed in almost all cells or tissue types (such as GAPDH); these are referred to as housekeeping genes. They are involved in functions common to all cells, such as basic energy metabolism, cell structure, etc. The relative amounts of RNA in the different lanes can be directly compared to see, e.g., which tissues express the gene most abundantly.
One can hybridize a labeled DNA clone to a panel of RNA samples from a progressive stages of development to determine the developmental stage when during development a particular cloned gene is expressed as RNA (bottom panel of Fig. 3.33). For instance, a gene product may be required for determination decisions early in development, and only be expressed in early embryos.
Once the DNA sequence of the gene of interest is known, and its intron-exon structure determined, highly sensitive RT-PCR assays can be designed (Fig. 3.34). The RNA from the cell or tissue of interest is copied into cDNA using reverse transcriptase and dNTPs, and then primers are annealed for PCR. Ideally, the primers are in different exons so that the product of amplifying the cDNA will be smaller than the product of amplifying the genomic DNA.

Figure 3.33.
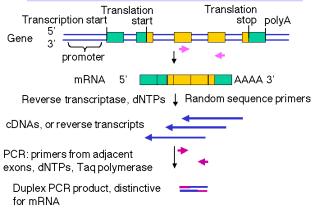
Figure 3.34. Reverse
transcription-PCR (RT-PCR) assay for mRNA.
In situ hybridizations / immunochemistry
In complementary approaches, the labeled DNA can be hybridized in situ to thin sections of a tissue or embryo or other specimen, and the resulting pattern of grains visualized along the specimen in the microscope (Fig. 3.35). Also, antibody probes against the protein product can be used to localize it in the specimen. This gives a more detailed picture of the pattern of expression, with resolution to the particular cells that are expressing the gene. The RNA blot-hybridization techniques described in a. above look at the RNA in all the cells from a tissue, and do not provide the level of resolution to single cells.
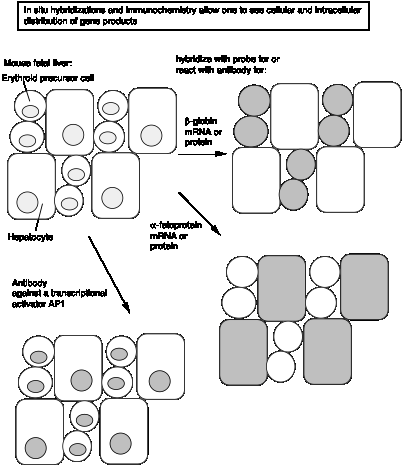
Figure 3.35.
Microarrays
As
large numbers of sequenced mRNAs and genes become available, technology has
been developed to look at expression of very large numbers of genes
simulatneously. DNA sequences specific for each gene in a bacterium or yeast
can be spotted in a high-density array with 400 r more spots. Some technologies
use many more spots, with mutliple sequences per gene. Microarrays, or “gene
chips” are available for many species, some with tens of thousands of different
sequences or “probes.” RNA from different tissues can be converted to cDNA with
a distinctive fluorescent label, and then hybridized to the gene chip.
Differences in level of expression
can be measured. Thus global changes in gene expression can now be measured.
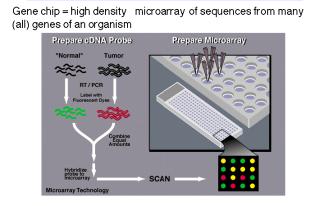
Figure 3.36. Hybridization of RNA to high density microarrays of gene sequences, or “gene chips”.
Database searches
An increasingly powerful approach is to determine candidates for the the function of your gene by searching the databases with the sequence, looking for matches to known proteins and genes. These matches provide clues as to protein function.
The power of this approach increases as the amount of sequences deposited in databases expand. Sequences of many genes are already known. The sequenced genes from more complex organisms, such as plants and animals, tend to be the ones more easily isolated using the techniques discussed in recombinant DNA technology. However, the sequences of genes expressed at a low level are starting to accumulate in the databases.
One remarkable advance in the past few years is the increasing number of organisms whose entire genome has been sequenced. About 10 bacterial genomes have been sequenced, and the number increases every few months. Genomics sequences for two eukaryotes are now available. That of the yeast Saccharomyces cerevisiae has been known for a few years, and the genome of the nematode Caenorhabditis elegans was completed in 1998. These sequences are being analyzed intensively, and a very high fraction of all the genes in each genome can be reliably detected using computational tools (one part of bioinformatics). It has become clear that many of the enzymes used in basic metabolism, regulation of the cell cycle, cellular signaling cascades, etc. are highly conserved across a broad phylogenetic spectrum. Thus it is common to find significant sequence matches in the genomes of model organisms when they are queried by the sequence of a previously unknown gene, e.g. from humans or mouse. The function already established for that gene in worms or yeast is a highly reliable guide to the function of the homologous gene in humans. The worm C. elegans is multicellular, and fate of each of its cells during development has been mapped. Thus it is possible that many functions involved in cellular interactions and cell-cell signaling will be conserved in this species, thus expanding the list of potential targets for a search in the databases.
This potential is being realized as working draft sequences of the human and mouse genomes are being analyzed. Within these data is a good approximation of sequences from virtually all human and mouse genes. Random clones have been partially sequenced from libraries of cDNAs from various human tissues, normalized to remove much of the products of abundant mRNAs and thus increasing the frequency of products of rare mRNAs. These sequences from the ends of the cDNA clones are called expressed sequence tags, or ESTs. The name is derived from the fact that since they are in cDNA libraries, they are obviously expressed at the level of mRNA, and some are used as tags in generating high-resolution maps of human chromosome. Hundreds of thousands of these have now been sequenced in collaborative efforts between pharmaceutical companies, other companies and universities. The database dbEST records all those in the public domain, and it is a strong complement to the databases recording all known sequences of genes. Many different parts of the same, or highly related, cDNAs, are recorded as separate entries in dbEST. Projects are underway to group all the sequences from the same (or highly related) gene into a a unified sequence. One example is the Unigene project at NCBI. The number of entries grows continually, but in the summer of 1998 there are about 50,000 entries, each representing about one gene. The number is higher now. Current estimates of the number of human genes are around 30,000, so it is possible that some UniGene clusters represent only parts of genes, and some genes match more than one cluster.
Very efficient search engines have been designed for handling queries to these databases, and several are freely available over the World Wide Web. One of the most popular and useful sites for this and related activities is maintained by the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/). Their Entrez browser provides integrated access to sequence, mapping and some functional information, PubMed provides access to abstracts of papers in journals in the National Library of Medicine, and the BLAST server allows rapid searches through various sequence databases. dbEST and the Unigene collection are maintained here, many genome maps are available, and three-dimensional structures of proteins and nucleic acids are available.
Make the protein product and analyze it
It is often possible to express the gene and make the encoded protein in large amounts. The protein can be purified and assayed for various enzymatic or other activities. Hypotheses for such activities may come from database searches.
Directed mutation
The previously describe approaches give some idea about gene function, but they do not firmly establish those functions. Indeed, this is a modern problem of trying to assign a function to an isolated gene. Several “reverse genetic” approaches can now be taken to tackle this problem. The most powerful approach to determining the physiological role(s) of a gene product is to mutate the gene in an appropriate organism and search for an altered phenotype.
The easiest experiment to do, but sometimes most difficult to interpret, is a gain of function assay. In this case, one forces expression of the gene in a transgenic organism, which often already has a wild type copy of the gene. One can look for a phenotype resulting from over-expression in tissues where it is normally expressed, or ectopic expression in tissues where it is normally silent.
In some organisms, it is possible to engineer a loss of function of the gene. The most effective method is to use homologous recombination to replace the wild type gene with one engineered to have no function. This knock-out mutation will prevent expression of the endogenous gene and one can see the effects on the whole organism. Unfortunately, the efficiency of homologous recombination is low in many organisms and cell lines, so this is not always feasible. Other methods for knocking out expression are being developed, although the mechanism for their effect (when successful) is still being studied. In some cases, one can block expression of the endogenous gene by forcing production of antisense RNA. Another method that is effective in some, but currently not all organisms, is the use of double-stranded, interfering RNA (RNAi). Duplex RNAs less than 30 nucleotide pairs long from the gene of interest can prevent expression of genes in worms, flies, and plants. Some success in mammals was recently reported.
Another way to generate a loss-of-function phenotype is to express dominant negative alleles of the gene. These mutant alleles encode stable proteins that form an aberrant structure that prevents functioning of the endogenous protein. This usually requires some protein-protein interaction (e.g. homodimers or heterodimers).
Localization on a genetic map
Sometimes the gene you have isolated maps to a region on a chromosome with a known function. Of course, many genes are probably located in that region, so it is critical to show that a candidate gene really is the one that when mutated causes an altered phenotype. This can be done by showing that a wild type copy of the candidate gene will restore a normal phenotype to the mutant. If a marker is known to be very tightly linked to the candidate gene, one can test whether this marker is always in linkage disequilibrium with the determinant of the mutant phenotype, i.e. in a large number of crosses, the marker for the candidate gene and the mutant phenotype never separated by recombination.
The mapping is often done with gene‑specific probes for in situ hybridizations to mitotic chromosomes. One then aligns the hybridization pattern with the chromosome banding patterns to map the isolated gene. Another method is to hybridize to a panel of DNAs from hybrid cells that contain only part of the chromosomal complement of the genome of interest. This is particularly powerful with radiation hybrid panels.
QUESTIONS
CHAPTER 3
ISOLATION AND
ANALYSIS OF GENES
3.2 Altering the ends of DNA fragments for ligation into vectors.
(Adapted from POB)
a) Draw the structure of the end of a linear DNA fragment that was generated by digesting with the restriction endonuclease EcoRI. Include those sequences remaining from the EcoRI recognition sequence.
b) Draw the structure resulting from the reaction of this end sequence with DNA polymerase I and the four deoxynucleoside triphosphates.
c) Draw the sequence produced at the junction if two ends with the structure derived in (b) are ligated.
d) Design two different short synthetic DNA fragments that would permit ligation of structure (a) with a DNA fragment produced by a PstI restriction digest. In one of these synthetic fragments, design the sequence so that the final junction contains the recognition sequences for both EcoRI and PstI. Design the sequence of the other fragment so that neither the EcoRI nor the PstI sequence appears in the junction.
3.3. What properties are required of vectors used in molecular cloning of DNA?
3.4. A student ligated a BamHI fragment containing a gene of interest to a pUC vector digested with BamHI, transformed E. coli with the mixture of ligation products and plated the cells on plates containing the antibiotic ampicillin and the chromogenic substrate X‑gal. Which colonies should the student pick to find the ones containing the recombinant plasmid (with the gene of interest in pUC)?
3.5. Starting with an isolated mRNA, one wishes to make a double stranded copy of the mRNA and insert it at the PstI site of pBR322 via G-C homopolymer tailing. One then transforms E. coli with this recombinant plasmid, selecting for tetracycline resistance. What are the four enzymatic steps used in preparing the cDNA insert? Name the enzymes and describe the intermediates.
3.6 A researcher needs to isolate a cDNA clone of giraffe actin mRNA, and she knows the size (Mr = 42,000) and partial amino acid sequence of giraffe actin protein and has specific antibodies against giraffe actin. After constructing a bank of cDNA plasmids from total mRNA of giraffe fibroblasts (dG-dC tailed into the PstI site of pBR322), what methods of screening the bank could be used to identify the actin cDNA clone?
3.7 The restriction map of pBR322 is
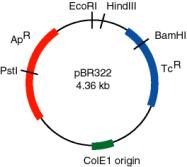
The distance in base pairs between restriction sites is as follows:
PstI to EcoRI 750 bp
EcoRI to HindIII 50 bp
HindIII to BamHI 260 bp
BamHI to PstI 3300 bp
A recombinant cDNA plasmid, pAlc-1, has double-stranded cDNA inserted at the PstI site of pBR322, using a technique that retains this cleavage site at both ends of the insert. Digestion of pBR322 and pAlc-1 with restriction endonucleases gives the following pattern after gel electrophoresis (left). The sizes of the fragments are given in base pairs. The DNA fragments were transferred out of the gel onto nitrocellulose and hybridized with radiolabeled cDNA from wild-type A. latrobus (a Southern blot-hybridizaton). Hybridizing fragments are shown in the autoradiogam diagram on the right.
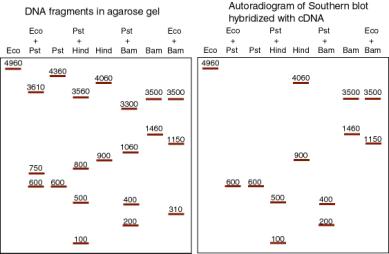

a) What is the size of the cDNA insert?
b) What two restriction endonucleases cleave within the cDNA insert?
c) For those two restriction endonucleases, each DNA fragment in the single digest is cut by PstI into two DNA fragments in the double digest (i.e. the restriction endonuclease plus PstI). Determine which fragments each single digest fragment is cut into, and use this information to construct a map.
d) Draw a restriction map for pAlc-1, showing sites for PstI, EcoRI, BamHI and HindIII. Indicate the distance between sites and show the cDNA insert clearly.
3.8. You isolate and clone a KpnI fragment from A. latrobus genomic DNA that encodes the mRNA cloned in pAlc-1 (as analyzed in question 3.7). The restriction map of the genomic fragment is

Each fragment that hybridizes to pAlc-1 is indicated by an asterisk. What does this map, especially when compared to that in problem 3.7, tell you about the structure of the gene? Be as quantitative as possible.
3.9. Some particular enzyme is composed of a polypeptide chain of 192 amino acids. The gene that encodes it has 1,440 nucleotide pairs. Explain the relationship between the number of amino acids in this polypeptide and the number of nucleotide pairs in its gene.
3.10. When viewed in the electron microscope, a hybrid between a cloned giraffe actin gene (genomic DNA) and mature actin mRNA looks like this:
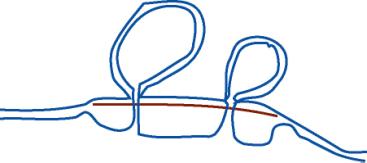

What can you conclude about actin gene structure in the giraffe?
3.11. DNA complementary to pepper mRNA was synthesized using oligo (dT) as a primer for first strand synthesis. The second strand (synonymous with the mRNA) was then synthesized, and the population of double stranded cDNAs were ligated into a plasmid vector using a procedure that leaves PstI sites flanking the cDNA insert (i.e. the terminal PstI sites for each clone are not part of the cDNA). This cDNA library was screened for clones made from the mRNA from the pepper yellow gene. One clone was isolated, and subsequent analysis of the pattern of restriction endonuclease cleavage patterns showed it had the following structure:

The map shows the positions of restriction endonuclease cleavage sites and the distance between them in kilobases (kb). The map of the cDNA insert is shown with solid lines, and plasmid vector DNA flanking the cDNA is shown as dotted lines. The top strand is oriented 5' to 3' from left to right, and the bottom strand is oriented 5' to 3' from right to left. The positions and orientations of two oligonucleotides to prime synthesis for sequence determination are shown, and are placed adjacent to the strand that will be synthesized in the sequencing reaction.
a) Oligonucleotides that anneal to the plasmid vector sequences that flank the duplex cDNA insert were used to prime synthesis of DNA for sequencing by the Sanger dideoxynucleotide procedure. A primer that annealed to the vector sequences to the left of the map shown above generated the sequencing gel pattern shown below on the left. A primer that annealed to the vector sequences to the right of the map shown above generated the sequencing gel pattern shown below on the right. The gels were run from the negative electrode at the top to the positive electrode at the bottom, and the segment presented is past the PstI site (i.e. do not look for a PstI recognition site).
Left primer Right primer
|
G |
A |
T |
C |
|
|
G |
A |
T |
C |
|
|
|
____ |
|
|
|
|
____ |
|
|
|
|
____ |
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
|
____ |
|
|
|
|
|
|
____ |
|
|
|
|
____ |
|
|
|
|
|
|
____ |
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
____ |
|
|
|
|
|
|
____ |
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
____ |
|
|
|
|
|
|
____ |
|
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
|
____ |
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
|
____ |
|
|
|
|
|
|
|
____ |
|
a) What is the DNA sequence of the left and right ends of the insert in the cDNA clone? Be sure to specify the 5' to 3' orientation, and the strand (top or bottom) whose sequence is reported. The terms left, right, top and bottom all refer to the map shown above for the cDNA clone.
b) Which end of the cDNA clone (left or right in the map above) is most likely to include the sequence synonymous with the 3' end of the mRNA?
c) What restriction endonuclease cleavage sites do you see in the sequencing data given?
3.12. Genomic DNA from the pepper plant was ligated into EcoRI sites in a l phage vector to construct a genomic DNA library. This library was screened by hybridization to the yellow cDNA clone. The pattern of EcoRI cleavage sites for one clone that hybridized to the yellow cDNA clone was analyzed in two experiments.
In the first experiment, the genomic DNA clone was digested to completion with EcoRI, the fragments separated on an agarose gel, transferred to a nylon filter, and hybridized with the radioactive yellow cDNA clone. The digest pattern (observed on the agarose gel) is shown in lane 1, and the pattern of hybridizing fragments (observed on an autoradiogram after hybridization) is shown in lane 2. Sizes of the EcoRI fragments are indicated in kb. The right arm of this l vector is 6 kb long, and the left arm is 30 kb.

In the second experiment, the genomic DNA clone was digested with a range of concentrations of EcoRI, so that the products ranged from a partial digest to a complete digest. The cleavage products were annealed to a radioactive oligonucleotide that hybridized only to the right cohesive end (cos site) of the l vector DNA. This simply places a radioactive tag at the right end of all the products of the reaction that extend to the right end of the l clone (partial or complete); digestion products that do not include the right end of the l clone will not be seen. The results of the digestion are shown above, on the right. Lane 1 is the clone of genomic DNA in l that has not been digested, lane 5 is the complete digest with EcoRI, and lanes 2, 3 and 4 are partial digests using increasing amounts of EcoRI. The sizes of the radioactive DNA fragments (in kb) are given, and the density of the fill in the boxes is proportional to the intensity of the signal on the autoradiogram.
a) What is the map of the EcoRI fragments in the genomic DNA clone, and which fragments encode mRNA for the yellow gene? You may wish to fill in the figure below; the left and right arms of the l vector are given. Show positions of the EcoRI cleavage sites, distances between them (in kb) and indicate the fragments that hybridize to the cDNA clone.
EcoRI EcoRI
(30 kb) (6 kb)
In a third experiment, the pepper DNA from the genomic DNA clone was excised, hybridized with yellow mRNA under conditions that favor RNA-DNA duplexes and examined in the electron microscope to visualize R-loops. A pattern like the following was observed. The lines in the figure can be duplex DNA, RNA-DNA duplexes and single-stranded DNA.

b) What do the R-loop data indicate? Please draw an interpretation of the R-loops, showing clearly the two DNA strands and the mRNA and distinguishing between the template (bottom, or message complementary) and nontemplate (top, or message synonymous) strands.
The EcoRI fragments that hybridize to the yellow cDNA clone were isolated and digested with SalI (S in the figure below), HindIII (H), and the combination of SalI plus HindIII (S+H). The resulting patterns of DNA fragments are shown below; all will hybridize to the yellow cDNA clone. Cleavage of the 5 kb EcoRI fragment with SalI generates two fragments of 2.5 kb.

c) What are the maps of the SalI and HindIII site(s) in each of the EcoRI fragments? Show positions of the cleavage sites and distances between them on the diagram below.
5 kb EcoRI fragment: 4 kb EcoRI fragment:
EcoRI EcoRI EcoRI EcoRI
|___________________________| |___________________________|
d) Compare these restriction maps with that of the cDNA clone (problem 1.38) and the R-loops shown above. Assuming that the SalI and HindIII sites in the genomic DNA correspond to those in the cDNA clone, what can you deduce about the intron/exon structure of the yellow gene(s) contained within the 5 kb and 4 kb EcoRI fragments? Please diagram the exon-intron structure in as much detail as the data permit (i.e. show the size of the intron(s) and positions of intron/exon junctions as precisely as possible).
5 kb EcoRI fragment: 4 kb EcoRI fragment:
EcoRI EcoRI EcoRI EcoRI
|___________________________| |___________________________|
e) Considering all the data (maps of cDNA and genomic clones and R-loop analysis), what can you conclude about the number and location(s) of yellow gene(s) in this genomic clone?
3.13 You have isolated an 1100 base pair (bp) cDNA clone for a gene called azure that when mutated causes blue eyes in frogs. You also isolate a 3000 bp SalI genomic DNA fragment that hybridizes to the azure cDNA. The map of the azure cDNA is as follows, with sizes of fragments given in bp.
![]()
Digestion of the 3000 bp SalI fragment of genomic DNA with the indicated restriction endonucleases yields the following pattern of fragments, all of which hybridize to the azure cDNA. Remember that the starting fragment has SalI sites at each end. Sizes of fragments are in bp.
Restriction enzymes
BamHI Bam+Pst PstI Pst+Eco EcoRI Bam+Eco
2700
2300
2000
1900 1900
1200
1100 1100
800
700 700 700
300 300 300
The SalI to SalI (3000 bp) genomic fragment was hybridized to the 1100 bp cDNA fragment, and the heteroduplexes were examined in the electron microscope. Measurements on a large number of molecules resulted in the determination of the sizes indicated in the structure on the left, i.e. duplex regions of 400 and 600 bp are interrupted by a single stranded loop of 1500 nucleotides and are flanked by single stranded regions of 500 and 100 nucleotides. When the same experiment is carried out with the 2700 bp SalI to EcoRI genomic DNA fragment hybridized to the cDNA fragment, the structure on the right is observed.
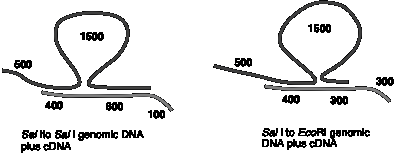
a) What is the restriction map of the 3000 bp SalI to SalI genomic DNA fragment from the azure gene? Specify distances between sites in base pairs.
b) How many introns are present in the azure genomic DNA fragment?
c) Where are the exons in the azure genomic DNA fragment? Draw the exons as boxes on the restriction map of the 3000 bp SalI to SalI genomic DNA fragment? Specify (in base pairs) the distances between restriction sites and the intron/exon boundaries.
3.14 The T-cell receptor is present only on T-lymphocytes, not on B-lymphocytes or other cells. Describe a strategy to isolate the T-cell receptor by subtractive hybridization, using RNA from T-lymphocytes and from B-lymphocytes.
3.15. How many exons are in the human insulin (INS) gene, how big are they, and how large are the introns that separate them? Use three different bioinformatic approaches to answer this.
a. Align the available genomic sequence containing INS (encoding insulin) with the sequence of the mRNA to find exons and introns in the INS gene. The sequence files are:
INS mRNA: accession number NM_000207
INS gene (includes part of TH and IGF2 in addition to INS): accession number L15440
Files can be obtained from NCBI (http://www.ncbi.nlm.nih.gov), or from the course web site (http://www.bmb.psu.edu/Courses/bmb400/default.htm)
Align the mRNA (cDNA) and genomic sequence using the BLAST2 sequences server at
http://www.ncbi.nlm.nih.gov/blast/
and the sim4 server at
Sim4 is designed to take into account terminal redundancy at the exon/intron junctions, whereas BLAST2 does not. Do you see this effect in the output?
b. Use the ab initio exon finding program Genscan, available at
http://genes.mit.edu/GENSCAN.html
to predict exons in the INS genomic sequence (L15440).
How does this compare with the results of analyzing with the program genscan?
c. What do you see for INS at the Human Genome Browser and Ensembl? They are accessed at:
http://genome.ucsc.edu/goldenPath/hgTracks.html
http://www.ensembl.org/