ANSWERS
to Questions from Part Three
Answers, Chapter 10. Transcription: RNA polymerases
10.1 The sigma factor (s) causes RNA polymerase to bind to the correct sites on DNA to initiate transcription (i.e. promoters). s destabilizes the complex between core polymerase and non-promoter DNA and decreases the amount of time it is bound. It enhances the affinity and increases the amount of time that holoenzyme (a2bb's) is bound to promoter, i.e. it facilitates a random search for promoters.
10.2 Statements 2 and 4 are correct.
10.3 Elongation of transcription by E. coli RNA polymerase proceeds at about 50 nucleotides per sec. Therefore, the rRNA primary transcript would be synthesized in 6500 nucleotides /50 nucleotides per sec = 130 sec, or slightly over 2 min.
10.4 0.83 initiations per sec. (50 nt/sec)(3.4 Angstroms/nt) = 170A/sec. 204A/170A sec-1 = 1.2 sec per initiation, or 0.83 initiations per sec.
10.5 a) True
b) False
c) True
d) True
10.6 Common features include:
a. All are template directed, synthesizing a sequence complementary to the template.
b. Synthesis occurs in a 5' to 3' direction.
c. All catalyze the addition of a nucleotide via the formation of a phosphodiester bond.
d. All release pyrophosphate as a product.
Distinctive features include:
a. The substrates: DNA polymerase, reverse transcriptase, and telomerase use deoxyribonucleoside triphosphates as a substrate, whereas RNA polymerase uses ribonucleoside triphosphates.
b. The templates: DNA polymerase and RNA polymerase use DNA as a template, whereas telomerase copies an RNA template that is part of the enzyme. Reverse transcriptase uses RNA as a template in the life cycle of retroviruses and retrotransposons, but in vitro it can use either DNA or RNA as a template.
c. Primer requirements: DNA polymerase, reverse transcriptase and telomerase require primers provided by some other activity or protein (primase, an tRNA or the 3’ end of a DNA strand, respectively), whereas RNA polymerase can begin synthesis of RNA internally to the template without a primer.
In general, the chemistry of the enzyme reaction is similar for all four, but the specific substrates, templates and primers differ.
Answers, Chapter 11. Transcription: Promoters and Terminators
11.1 a) Right to left
b) 1800
c) 400
5' end label: The lack of protection of the labeled Kpn-Bgl* 500 nucleotide fragment tells you that the mRNA is synonymous with the bottom strand, and thus the top strand is the template strand. The top strand is labeled at the 5' end of the 1500 nucleotide *Bgl-Kpn fragment, and hybridization of this probe with mRNA gives protection of a 1300 nt fragment. This indicates that transcription proceeds from right to left (on the map as given), and the 5'' end of the transcript is 1300 nts to the right of the BglII site. In the coordinates of the map, this would be 500 (position of BglII) + 1300 = 1800.
5'Kpn 500 Bgl Bgl 1500 Kpn
---------------------> *--------------------------------------->
<--------------------* <---------------------------------------
ß ¯
no protection
5' Bgl Kpn
*--------------------------------------->
| | | | | | | | | | | | | | | | | | | | | | | | |
<~~~~~~~~~~~~~~~~~~~~~~~~~•cap
¯ S1
¯ gel
*--------------------------->
1300 nt protected fragment
When the 5' end label is at the EcoRI site, a similar result is obtained, but one can map the 5' end with greater accuracy. The protected fragment is from the top strand and is 100 nts (a size that can be measured more accurately than the 1300 nt fragment on the polyacrylamide gels used in this analysis).
Kpn 1700 Eco Eco 300 Kpn
*--------------------------------------------> *----------------->
<--------------------------------------------* ------------------
¯
*----------------->
| | | | |
<~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~•
¯ S1,..etc.
100 nt
*------->
3' end label: When the DNA fragments are labeled at the 3' end, again the top strand will be protected by hybridization to mRNA, thus reaffirming the conclusions above that the top strand is the template strand. The 500 nt Kpn-Bgl* fragment generates a 100 nt protected fragment, showing that the 3' end of the mRNA is 100 nts to the left of the BglII site, or at 500 - 100 = 400 on the coordinates of the map.
5'Kpn 500 3'Bgl Bgl 1500 Kpn
----------------->* ------------------------------------>
----------------- *------------------------------------
¯ ß
5' -----------------* no protection
| | | |
<~~~~~~~~~~~~~~~~~~~~•
¯ S1, gel,..etc.
-----*
100 nt
11.2 a) Left to right.
b) 400
c) Cannot be determined.
11.3 a) It will increase expression of the almond gene.
b) It has no effect.
c) It will increase expression of the almond gene.
d) The -50 to -1 fragment is acting like a promoter. In the first set of experiments, it is needed for promotion of transcription and it is needed to respond to upstream activating sequences. In the second set of experiments, the heterologous promoter will substitute for it.
11.4 a) Two complexes are formed between the labeled probe and the kidney cell nuclear extract.
b) Lanes 3-8 tell you that complexes A and B are specific, i.e. the proteins are recognizing a particular DNA sequence, since the self-DNA competes, but the E. coli DNA does not.
c) Lanes 9-14 tell you that the protein binding to form complex A will also bind to a DNA containing an Sp1 binding site. Thus the protein that forms complex A with this probe may be Sp1 or a relative of this protein. Neither protein (for complex A or complex B) will bind to the DNA probe with the Oct1 binding site, showing that this is not a candidate for the protein forming the sequence-specific complexes with the probe.
11.5 5' GAGTC
3' CTCAG
11.6 a) False
b) True
c) True
d) True
Answers to questions in Chapter 12. RNA Processing
12.1 a) NTP labeled at either the b or the g position.
b) NTP labeled at the b position, which will be in the cap structure.
c) NTP labeled at the a position, since the b and g phosphates are lost upon incorporation of the NMP.
12.2 One of the key signals for cleavage and 3' polyadenylation is the sequence AAUAAA. After RNA polymerase II has transcribed beyond this sequence, an endonuclease (uncharacterized at this time) cleaves the primary transcript at a position about 25 to 30 nucleotides 3' to the AAUAAA. Then the enzyme polyadenylate polymerase adds a string of 20 to 250 A's to the free 3' end, generating the 3' poly(A) tail.
The mutation would prevent cleavage and polyadenylation at the usual site, which could have two different consequences. If the transcript is not polyadenylated, it will be quite unstable and the steady state levels of mRNA will be very low, and therefore little or no protein product will be made. In some cases, a "substitute AAUAAA" may happen to occur in the transcribed region 3' to the gene, in which case this alternative, "cryptic" polyadenylation site could be used. However, this cryptic site may not be used as efficiently as the wild type (normal) sequence, again resulting in a reduction in the amount of steady state mRNA.
12.3 a) False
b) True
c) True
d) True
12.4 Both are capable of self-splicing, and both utilize a phosphoester transfer mechanism (transesterification) that is initiated by a guanine nucleoside or nucleotide. Neither require ATP hydrolysis.
12.5 a) Introns almost invariably have the dinucleotides GU at their 5' end and AG at their 3' end.
b) U1 snRNP
c) A branch structure forms by linkage between the 2' hydroxyl of an A at the branch site in the intron and the 5' phosphate at the 5' G of the intron.
d) Spliceosome assembly require hydrolysis of ATP.
12.6 The mechanism for removal of introns from pre‑mRNAs is very similar to that of the Group II introns, with the formation of a lariat intermediate after the reaction is initiated. Each of the cleavage and rejoining reactions is a transesterification, in which a new phosphodiester bond is formed for every one that is broken.
A minimum of two transesterification steps are required. The first step is initiated by the attack of a 2' hydroxyl of an A within the intron on the bond linking the 3' end of the first exon with the 5' end of the intron. This generates a 3' hydroxyl on the nucleotide at the 3' end of the first exon, and effectively takes the intron out of the series of transesterifications by forming a lariat structure. This 3' nucleotide of the first intron can then link to the first nucleotide of the second exon, again by a transesterification. The result of this second step is the union of the first and second exons, with the intron liberated as a lariat intermediate.
12.7 a) 2, 5
b) 3
c) 1, 4
12.8 (a) The excised intron has a branch point where an RNA chain is covalently attached to the original chain of RNA.
(b) The excised intron has a circle and a linear tail, i.e. it is a lariat.
(c) Cleavage in the region complementary to oligo 1 will linearize the nonlinear structure and generate a molecule that is just about a full-length intron. E.g. it could open the circular part of a lariat, but it opens it adjacent to the branch or joint.
(d) The combined results with both oligos 3 and 4 show that branch point in the excised intron is located between the segments complementary to oligodeoxyribonucleotides 3 and 4.
(e) The structure of the excised intron is:
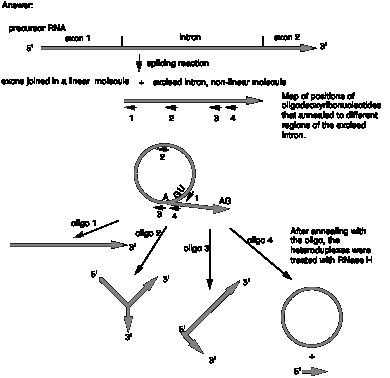
Answers
to questions from Chapter 13. Genetic Code
13.1 DNA and RNA polymerases have several properties in common.
1. All are template directed, synthesizing a sequence complementary to the template.
2. Synthesis occurs in a 5' to 3' direction.
3. All catalyze the addition of a nucleotide via the formation of a phosphodiester bond.
4. All use (deoxy)ribonucleoside triphosphates as a substrate, and release pyrophosphate as a product.
The enzyme polynucleotide phosphorylase can be used to synthesize RNA in vitro, and this was a key technique in deciphering the genetic code. However, it differs from DNA and RNA polymerases in points 1 and 4. Polynucleotide phosphorylase does not use a template, but rather adds ribonucleotides to an RNA in a highly reversible reaction. The substrates (in the direction of synthesis) are ribonucleoside diphosphates, which are added with the release of phosphate as a product. In the cell, this enzyme probably catalyzes the reverse reaction to degrade RNAs.
13.2 a) It starts at AUG (nucleotides 5-7) and ends at UGA (nucleotides 23-25). 6 amino acids are encoded, including the initiating methionine.
b) All codons have a U at the second position, hence only hydrophobic amino acids are encoded.
13.3 a) More than 1 codon encodes an amino acid.
b) 3rd.
c) The base in the 5' position of the anticodon can often pair with several bases in the "wobble" or 3rd position of the codon (e.g. I with C, U, or A). Therefore, one tRNA can recognize several codons.
13.4 The template strand is the strand that serves as the template for RNA synthesis; nontemplate strand is identical in sequence with the RNA transcribed from the gene, with U in place of T..
a) (5')CGACGGCGCGAAGUCAGGGGUGUUAAG(3')
b) Arg-Arg-Arg-Glu-Val-Arg-Gly-Val-Lys
c) No; The base sequence of mRNA transcribed from the nontemplate strand would be: (5')CUUAACACCCCTGACUUCGCGCCGUCG. This mRNA when translated would result in a different peptide than in (b). The complementary antiparallel strands in double-helical DNA do not have the same base sequence in the 5'to 3' direction. RNA is transcribed from only one specific strand of duplex DNA. The RNA polymerase must therefore recognize and bind to the correct strand.
13.5 The two DNA codons for Glu are GAA and GAG, and the four DNA codons for Val are GTT, GTC, GTA, and GTG. A single-base change in GAA to form GTA or in GAG to form GTG could account for the Glu Æ Val replacement in sickle-cell hemoglobin. Much less likely are two-base changes from GAA to GTG, GTT, or GTC; and from GAG to GTA, GTT, or GTC.
13.6 AAA
13.7 a) CAG
b) UUG
13.8 a) Three tRNAs are required. Anticodons 3' GCI and 3' GCC can accommodate the 5' CGN codons, and anticodon 3' UCU will pair with the 5' AGR codons.
b) Two tRNAs are required. The 5' GUN codon can be matched with anticodons 3' CAI + 3' CAC, or 3' CAG + 3' CAU.
13.9 a) Glycine should attach to a tRNAs with codons 5' GGU, 5' GGC and 5' GGA.
b) Isoleucine should attach to tRNAs with codons 5' AUC and 5' AUU.
13.10 Some amino acids are encoded by 6 different codons, some 4 different codons, some 3 different codons, some 2 different codons, and some one codon. To minimize the degree of ambiguity in codon assignment for a given peptide sequence, one must select a region of the peptide that contains mostly amino acids specified by a small number of codons.
Focus on the amino acids with the fewest codons: Met and Trp. The best possibility is the span of DNA from the codon for the first Trp residue to the first two nucleotides of the codon for Ile. The sequence of the probe would be:
(5')UGGUA(U/C)UG(U/C)AUGGA(U/C)UGGAU
The synthesis would be designed to incorporate either U or C where indicated, producing a mixture of eight 20-nucleotide probes that differ only at one or more of these positions.
13.11 a) 2 nucleotides
b) No
c) AA Codon
1 KK
2 NN
3 DD
4 KN
5 NK
6 DK
7 KD
8 ND
d) DN
e) DK ® NK
aa6 aa5
f) ND ® KD
aa8 aa7
Answers to Chapter 14. Translation
14.1 There are two tRNAs for methionine: tRNAfMet, the initiating tRNA, and tRNAMet, which can insert Met in interior positions in a polypeptide. tRNAfMet reacts with Met to yield Met-tRNAfMet, promoted by methionine aminoacyl-tRNA synthetase. The amino group of its Met residue is then formylated by N 10-formyltetrahydrofolate to yield fMet-tRNAfMet. Free Met or Met-tRNAMet cannot be formylated. Only fMet-tRNAfMet is recognized by the initiation factor IF-2 and is aligned with the initiating AUG positioned at the ribosomal P site in the initiation complex. AUG codons in the interior of the mRNA are eventually positioned at the ribosomal P site and can bind and incorporate only Met-tRNAMet.
14.2 a) True
b) False
c) True
d) True
14.3 a) The C terminal peptide.
b) All peptides have the same specific activity.
14.4 a) EF-G-GTP
b) IF-2, with GTP
c) RF-1
d) The 16S rRNA in the small ribosomal subunit.
14.5 a) Small or 30 S
b) AGGA in mRNA is complementary to the 3' end of 16S rRNA occurs before the AUG initiation codon.
c) IF3; IF2-f-Met-tRNA-GTP; IF1; 50S ribosomal subunit
14.6 1) aa + tRNA + ATP ® AMP + PPi + aa - tRNA
aa - tRNA synthetase
2) Binding of aa-tRNA to A site on ribosome;
requires EF-Tu and GTP ® GDP + Pi
3) Translocation of peptidyl-tRNA to P site on ribosome;
requires EF-G and GTP ® GDP + Pi
14.7 The amino acid most recently added to a growing polypeptide chain is the only one covalently attached to a tRNA and hence is the only link between the polypeptide and the mRNA that is encoding it. A proofreading activity would sever this link, halting synthesis of the polypeptide and releasing it from the mRNA.
14.8 An E. coli promoter is required for transcription, because E. coli RNA polymerase does not interact with eukaryotic promoters; a ribosome binding site positioned at an appropriate distance upstream from the ATG codon is required, because eukaryotic mRNA does not utilize such a site for translation initiation; an operator site is required for regulation of transcription in E. coli.
14.9 One hypothesis would be that the codon pair AUY has "always" encoded isoleucine, and in early evolution, the codon pair AUR encoded methionine. Subsequent specialization in the use of AUG has allowed it exclusively to be used to encode methionine, thereby allowing AUA to be recruited as an additional codon for isoleucine. This hypothesis assumes that both isoleucine and methionine were used in proteins early in evolution.
An alternative hypothesis states that AUN was originally a codon family encoding isoleucine and subseqently, the AUG was recruited to encode methionine. This hypothesis requires either that methionine was not used in proteins early in evolution, or that it was encoded by some other codon besides AUG. Neither of these latter possibilities seems very likely, so the alternative hypothesis is harder to rationalize. However, it cannot be ruled out, since we do not have direct access to observe the conditions of early evolution.
14.10 (a) 1 and 5 are correct.
(b) 2, 4, 7 are correct.
(c) 3 and 6 are correct.